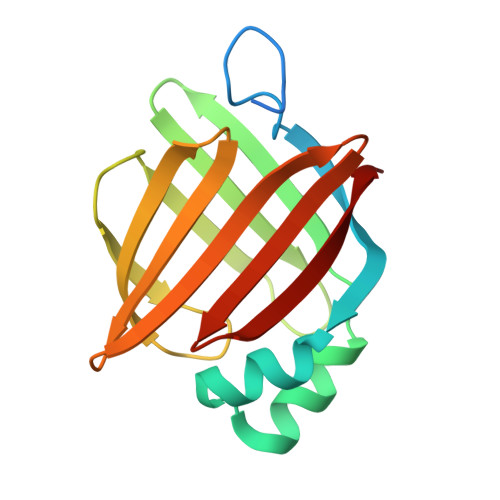
Structural Basis for Activation of Fatty Acid-binding Protein 4
Gillilan, R.E., Ayers, S.D., Noy, N.(2007) J Mol Biology 372: 1246-1260
- PubMed: 17761196 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2007.07.040
- Primary Citation Related Structures:
2Q9S, 2QM9 - PubMed Abstract:
Fatty acid-binding protein 4 (FABP4) delivers ligands from the cytosol to the nuclear receptor PPARgamma in the nucleus, thereby enhancing the transcriptional activity of the receptor. Notably, FABP4 binds multiple ligands with a similar affinity but its nuclear translocation is activated only by specific compounds. To gain insight into the structural features that underlie the ligand-specificity in activation of the nuclear import of FABP4, we solved the crystal structures of the protein complexed with two compounds that induce its nuclear translocation, and compared these to the apo-protein and to FABP4 structures bound to non-activating ligands. Examination of these structures indicates that activation coincides with closure of a portal loop phenylalanine side-chain, contraction of the binding pocket, a subtle shift in a helical domain containing the nuclear localization signal of the protein, and a resultant change in oligomeric state that exposes the nuclear localization signal to the solution. Comparisons of backbone displacements induced by activating ligands with a measure of mobility derived from translation, libration, screw (TLS) refinement, and with a composite of slowest normal modes of the apo state suggest that the helical motion associated with the activation of the protein is part of the repertoire of the equilibrium motions of the apo-protein, i.e. that ligand binding does not induce the activated configuration but serves to stabilize it. Nuclear import of FABP4 can thus be understood in terms of the pre-existing equilibrium hypothesis of ligand binding.
- Macromolecular Diffraction Facility of the Cornell High-Energy Synchrotron Source, Ithaca, NY 14853, USA.
Organizational Affiliation: