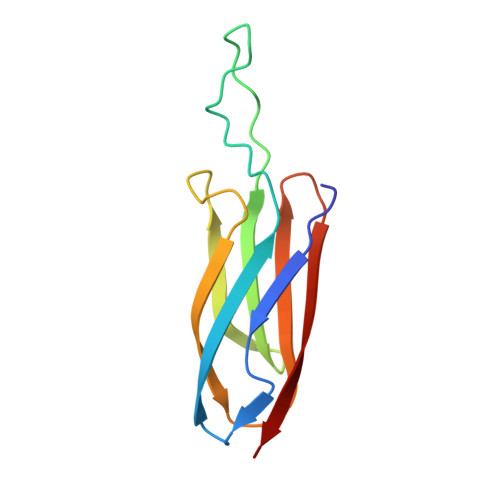
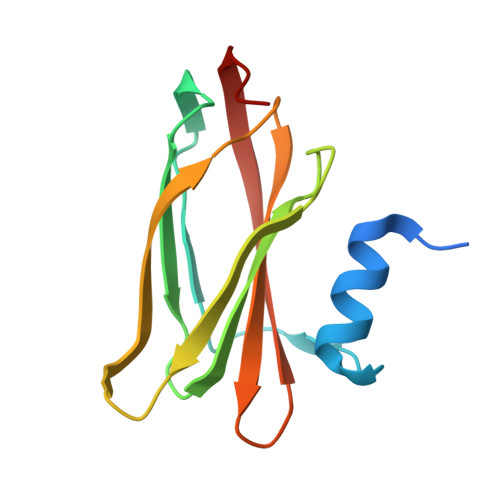
The SoxYZ Complex Carries Sulfur Cycle Intermediates on a Peptide Swinging Arm.
Sauve, V., Bruno, S., Berks, B.C., Hemmings, A.M.(2007) J Biological Chem 282: 23194-23204
- PubMed: 17522046 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M701602200
- Primary Citation Related Structures:
2OX5, 2OXG, 2OXH - PubMed Abstract:
The bacterial Sox (sulfur oxidizing) system allows the utilization of inorganic sulfur compounds in energy metabolism. Central to this process is the SoxYZ complex that carries the pathway intermediates on a cysteine residue near the C terminus of SoxY. Crystal structures have been determined for Paracoccus pantotrophus SoxYZ with the carrier cysteine in the underivatized state, conjugated to the polysulfide mimic beta-mercaptoethanol, and as the sulfonate adduct pathway intermediate. The carrier cysteine is located on a peptide swinging arm and is bracketed on either side by diglycine dipeptides acting as molecular universal joints. This structure provides a novel solution to the requirement that the cysteine-bound intermediates be able to access and orient themselves within the active sites of multiple partner enzymes. Adjacent to the swinging arm there is a conserved, deep, apolar pocket into which the beta-mercaptoethanol adduct extends. This pocket would be well suited to a role in protecting labile pathway intermediates from adventitious reactions.
- Department of Biochemistry, University of Oxford, Oxford OX1 3QU, United Kingdom.
Organizational Affiliation: