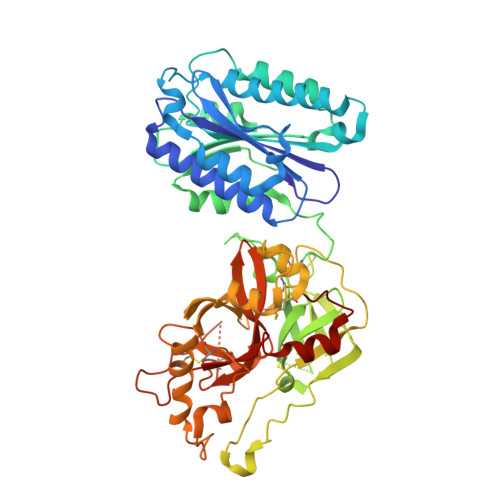
The crystal structure of c2a, the catalytic fragment of classical pathway c3 and c5 convertase of human complement.
Krishnan, V., Xu, Y., Macon, K., Volanakis, J.E., Narayana, S.V.(2007) J Mol Biology 367: 224-233
- PubMed: 17234210 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2006.12.039
- Primary Citation Related Structures:
2ODP, 2ODQ - PubMed Abstract:
The multi-domain serine protease C2 provides the catalytic activity for the C3 and C5- convertases of the classical and lectin pathways of complement activation. Formation of these convertases requires the Mg(2+)-dependent binding of C2 to C4b, and the subsequent cleavage of C2 by C1s or MASP2, respectively. The C-terminal fragment C2a consisting of a serine protease (SP) and a von Willebrand factor type A (vWFA) domain, remains attached to C4b, forming the C3 convertase, C4b2a. Here, we present the crystal structure of Mg(2+)-bound C2a to 1.9 A resolution in comparison to its homolog Bb, the catalytic subunit of the alternative pathway C3 convertase, C3bBb. Although the overall domain arrangement of C2a is similar to Bb, there are certain structural differences. Unexpectedly, the conformation of the metal ion-dependent adhesion site and the position of the alpha7 helix of the vWFA domain indicate a co-factor-bound or open conformation. The active site of the SP domain is in a zymogen-like inactive conformation. On the basis of these structural features, we suggest a model for the initial steps of C3 convertase assembly.
- Center for Biophysical Sciences and Engineering, School of Optometry, University of Alabama at Birmingham, Birmingham, AL 35294, USA.
Organizational Affiliation: