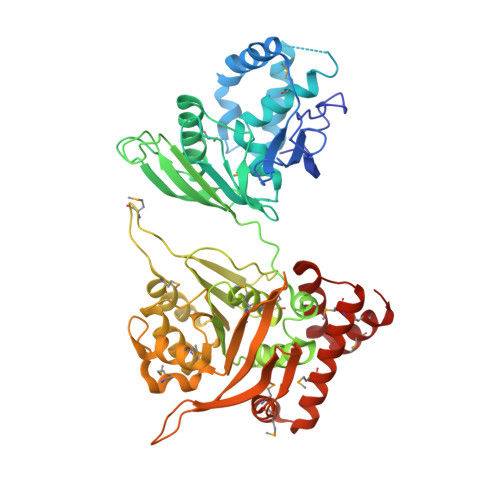
Hexameric structures of the archaeal secretion ATPase GspE and implications for a universal secretion mechanism.
Yamagata, A., Tainer, J.A.(2007) EMBO J 26: 878-890
- PubMed: 17255937 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601544
- Primary Citation Related Structures:
2OAP, 2OAQ - PubMed Abstract:
The secretion superfamily ATPases are conserved motors in key microbial membrane transport and filament assembly machineries, including bacterial type II and IV secretion, type IV pilus assembly, natural competence, and archaeal flagellae assembly. We report here crystal structures and small angle X-ray scattering (SAXS) solution analyses of the Archaeoglobus fulgidus secretion superfamily ATPase, afGspE. AfGspE structures in complex with ATP analogue AMP-PNP and Mg(2+) reveal for the first time, alternating open and closed subunit conformations within a hexameric ring. The closed-form active site with bound Mg(2+) evidently reveals the catalytically active conformation. Furthermore, nucleotide binding results and SAXS analyses of ADP, ATPgammaS, ADP-Vi, and AMP-PNP-bound states in solution showed that asymmetric assembly involves ADP binding, but clamped closed conformations depend on both ATP gamma-phosphate and Mg(2+) plus the conserved motifs, arginine fingers, and subdomains of the secretion ATPase superfamily. Moreover, protruding N-terminal domain shifts caused by the closed conformation suggest a unified piston-like, push-pull mechanism for ATP hydrolysis-dependent conformational changes, suitable to drive diverse microbial secretion and assembly processes by a universal mechanism.
- Department of Molecular Biology, The Skaggs Institute for Chemical Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: