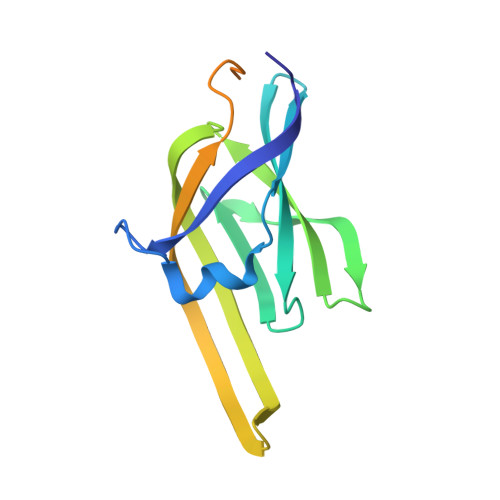
Crystal structure of the heme-IsdC complex, the central conduit of the Isd iron/heme uptake system in Staphylococcus aureus.
Sharp, K.H., Schneider, S., Cockayne, A., Paoli, M.(2007) J Biological Chem 282: 10625-10631
- PubMed: 17287214
- DOI: https://doi.org/10.1074/jbc.M700234200
- Primary Citation Related Structures:
2O6P - PubMed Abstract:
Pathogens such as Staphylococcus aureus require iron to survive and have evolved specialized proteins to steal heme from their host. IsdC is the central conduit of the Isd (iron-regulated surface determinant) multicomponent heme uptake machinery; staphylococcal cell-surface proteins such as IsdA, IsdB, and IsdH are thought to funnel their molecular cargo to IsdC, which then mediates the transfer of the iron-containing nutrient to the membrane translocation system IsdDEF. The structure of the heme-IsdC complex reveals a novel heme site within an immunoglobulin-like domain and sheds light on its binding mechanism. The folding topology is reminiscent of the architecture of cytochrome f, cellobiose dehydrogenase, and ethylbenzene dehydrogenase; in these three proteins, the heme is bound in an equivalent position, but interestingly, IsdC features a distinct binding pocket with the ligand located next to the hydrophobic core of the beta-sandwich. The iron is coordinated with a tyrosine surrounded by several non-polar side chains that cluster into a tightly packed proximal side. On the other hand, the distal side is relatively exposed with a short helical peptide segment that acts as a lip clasping onto almost half of the porphyrin plane. This structural feature is argued to play a role in the mechanism of binding and release by switching to an open conformation and thus loosening the interactions holding the heme. The structure of the heme-IsdC complex provides a template for the understanding of other proteins, such as IsdA, IsdB, and IsdH, that contain the same heme-binding module as IsdC, known as the NEAT (near transporter) domain.
- School of Pharmacy, Centre for Biomolecular Sciences, University of Nottingham, University Park, Nottingham NG7 2RD, United Kingdom.
Organizational Affiliation: