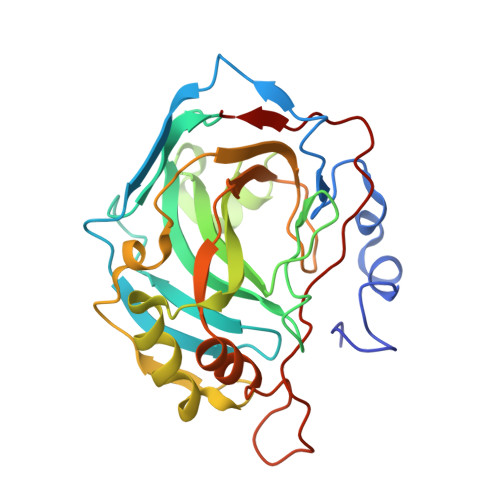
Carbonic anhydrase inhibitors: The X-ray crystal structure of the adduct of N-hydroxysulfamide with isozyme II explains why this new zinc binding function is effective in the design of potent inhibitors.
Temperini, C., Winum, J.Y., Montero, J.L., Scozzafava, A., Supuran, C.T.(2007) Bioorg Med Chem Lett 17: 2795-2801
- PubMed: 17346964
- DOI: https://doi.org/10.1016/j.bmcl.2007.02.068
- Primary Citation Related Structures:
2O4Z - PubMed Abstract:
N-Hydroxysulfamide is a 2000-fold more potent inhibitor of the zinc enzyme carbonic anhydrase (CA, EC 4.2.1.1) as compared to sulfamide. It also inhibits other physiologically relevant isoforms, such as the tumor-associated CA IX and XII (K(I)s in the range of 0.865-1.34microM). In order to understand the binding of this inhibitor to the enzyme active site, the X-ray crystal structure of the human hCA II-N-hydroxysulfamide adduct was resolved. The inhibitor coordinates to the active site zinc ion by the ionized primary amino group, participating in an extended network of hydrogen bonds with amino acid residues Thr199, Thr200 and two water molecules. The additional two hydrogen bonds in which N-hydroxysulfamide bound to hCA II is involved as compared to the corresponding adduct of sulfamide may explain its higher affinity for the enzyme, also providing hints for the design of tight-binding CA inhibitors possessing an organic moiety substituting the NH group in the N-hydroxysulfamide structure.
- Università degli Studi di Firenze, Polo Scientifico, Laboratorio di Chimica Bioinorganica, Rm. 188, Via della Lastruccia 3, 50019 Sesto Fiorentino (Florence), Italy. claudiu.supuran@unifi.it
Organizational Affiliation: