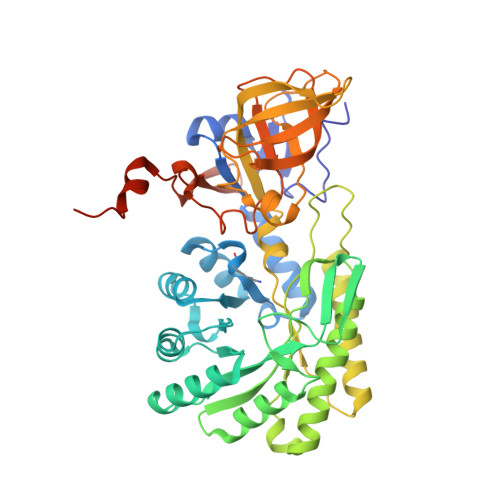
The three-dimensional structure of diaminopimelate decarboxylase from Mycobacterium tuberculosis reveals a tetrameric enzyme organisation.
Weyand, S., Kefala, G., Svergun, D.I., Weiss, M.S.(2009) J Struct Funct Genomics 10: 209-217
- PubMed: 19543810 Search on PubMed
- DOI: https://doi.org/10.1007/s10969-009-9065-z
- Primary Citation Related Structures:
2O0T - PubMed Abstract:
The three-dimensional structure of the enzyme diaminopimelate decarboxylase from Mycobacterium tuberculosis has been determined in a new crystal form and refined to a resolution of 2.33 A. The monoclinic crystals contain one tetramer exhibiting D(2)-symmetry in the asymmetric unit. The tetramer exhibits a donut-like structure with a hollow interior. All four active sites are accessible only from the interior of the tetrameric assembly. Small-angle X-ray scattering indicates that in solution the predominant oligomeric species of the protein is a dimer, but also that higher oligomers exist at higher protein concentrations. The observed scattering data are best explained by assuming a dimer-tetramer equilibrium with about 7% tetramers present in solution. Consequently, at the elevated protein concentrations in the crowded environment inside the cell the observed tetramer may constitute the biologically relevant functional unit of the enzyme.
- EMBL Hamburg Outstation, c/o DESY, Hamburg, Germany.
Organizational Affiliation: