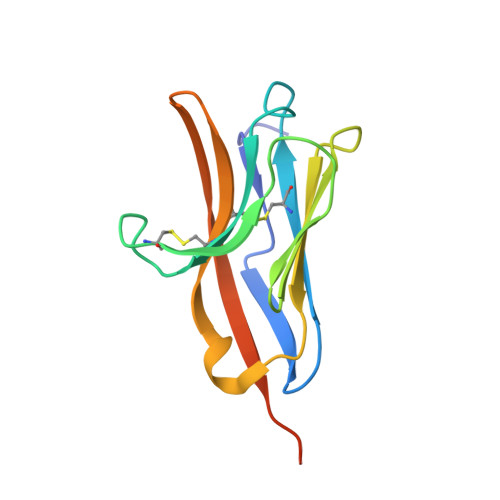
The Crystal Structure of the Extracellular Domain of the Inhibitor Receptor Expressed on Myeloid Cells IREM-1.
Marquez, J.A., Galfre, E., Dupeux, F., Flot, D., Moran, O., Dimasi, N.(2007) J Mol Biology 367: 310-318
- PubMed: 17275839
- DOI: https://doi.org/10.1016/j.jmb.2007.01.011
- Primary Citation Related Structures:
2NMS - PubMed Abstract:
The immune receptors expressed on myeloid cells (IREM) are type I transmembrane proteins encoded on human chromosome 17 (17q25.1), whose function is believed to be important in controlling inflammation. To date, three IREM receptors have been identified. IREM-1 functions as an inhibitory receptor, whereas IREM-2 and IREM-3 serve an activating function. Here, we report the crystal structure of IREM-1 extracellular domain at 2.6 A resolution. The overall fold of IREM-1 resembles that of a V-type immunoglobulin domain, and reveals overall close homology with immunoglobulin domains from other immunoreceptors such as CLM-1, TREM-1, TLT-1 and NKp44. Comparing the surface electrostatic potential and hydrophobicity of IREM-1 with its murine homologous CLM-1, we observed unique structural properties for the complementary determining region of IREM-1, which suggests that they may be involved in recognition of the IREM-1 ligand. Particularly interesting is the structural conformation and physical properties of the antibody's equivalent CDR3 loop, which we show to be a structurally variable region of the molecule and therefore could be the main structural determinant for ligand discrimination and binding. In addition, the analysis of the IREM-1 structure revealed the presence of four structurally different cavities. Three of these cavities form a continuous hydrophobic groove on the IREM-1 surface, which point to a region of the molecule capable of accommodating potential ligands.
- European Molecular Biology Laboratory, Grenoble Outstation Polygon Scientifique, 6 Rue Jules Horowitz, 38000 Grenoble, France.
Organizational Affiliation: