Solution Nuclear Magnetic Resonance Studies of Sterol Carrier Protein 2 Like 2 (SCP2L2) Reveal the Insecticide Specific Structural Characteristics of SCP2 Proteins in Aedes aegypti Mosquitoes
Singarapu, K.K., Ahuja, A., Potula, P.R., Ummanni, R.(2016) Biochemistry 55: 4919-4927
- PubMed: 27508310 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.6b00322
- Primary Citation Related Structures:
2NBM, 2NBN - PubMed Abstract:
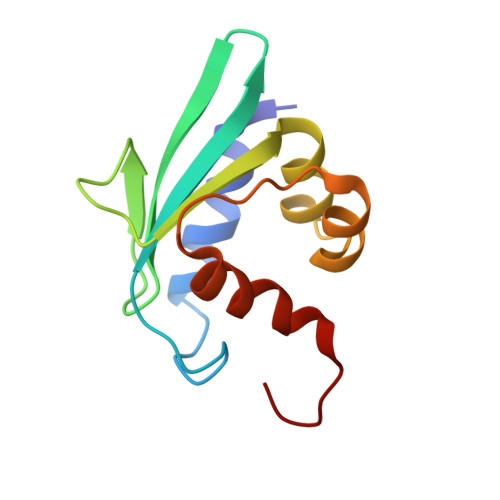
Sterol carrier protein 2 like 2 from Aedes aegypti (AeSCP2L2) plays an important role in lipid transport in mosquitoes for its routine metabolic processes. Repeated unsuccessful attempts to crystallize ligand free SCP2L2 prompted us to undertake nuclear magnetic resonance (NMR) spectroscopy to determine its three-dimensional structure. We report here the three-dimensional structures and dynamics of apo-AeSCP2L2 and its complex with palmitate. The (15)N heteronuclear single-quantum coherence spectrum of apo-AeSCP2L2 displayed multiple peaks for some of the amide resonances, implying the presence of multiple conformations in solution, which are transformed to a single conformation upon formation of the complex with plamitate. The three-dimensional structures of apo-AeSCP2L2 and palmitated AeSCP2L2 reveal an α/β mixed fold, with five β-strands and four α-helices, very similar to the other SCP2 protein structures. Unlike the crystal structure of palmitated AeSCP2L2, both solution structures are monomeric. It is further confirmed by the rotational correlation times determined by NMR relaxation times (T1 and T2) of the amide protons. In addition, the palmitated AeSCP2L2 structure contains two palmitate ligands, bound in the binding pocket, unlike the three palmitates bound in the dimeric form of AeSCP2L2 in the crystals. The relaxation experiments revealed that complex formation significantly reduces the dynamics of the protein in solution.
- Center for NMR and Structural Chemistry and §Center for Chemical Biology, CSIR-Indian Institute of Chemical Technology , Uppal Road, Tarnaka, Hyderabad 500007, India.
Organizational Affiliation: