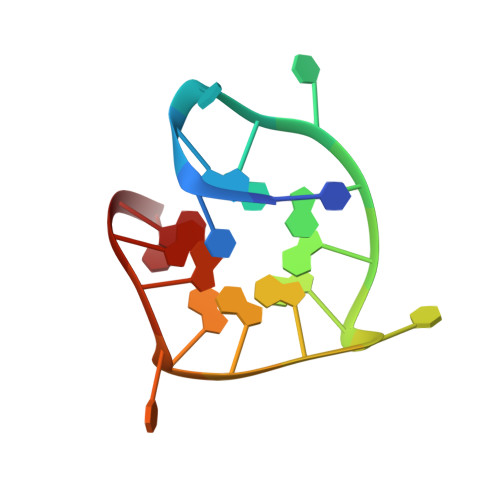
G-quadruplexes with (4n - 1) guanines in the G-tetrad core: formation of a G-triadwater complex and implication for small-molecule binding
Heddi, B., Martin-Pintado, N., Serimbetov, Z., Kari, T.M., Phan, A.T.(2016) Nucleic Acids Res 44: 910-916
- PubMed: 26673723 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkv1357
- Primary Citation Related Structures:
2N60 - PubMed Abstract:
G-quadruplexes are non-canonical structures of nucleic acids, in which guanine bases form planar G-tetrads (G·G·G·G) that stack on each other in the core of the structure. G-quadruplexes generally contain multiple times of four (4n) guanines in the core. Here, we study the structure of G-quadruplexes with only (4n - 1) guanines in the core. The solution structure of a DNA sequence containing 11 guanines showed the formation of a parallel G-quadruplex involving two G-tetrads and one G-triad with a vacant site. Molecular dynamics simulation established the formation of a stable G-triad·water complex, where water molecules mimic the position of the missing guanine in the vacant site. The concept of forming G-quadruplexes with missing guanines in the core broadens the current definition of G-quadruplex-forming sequences. The potential ability of such structures to bind different metabolites, including guanine, guanosine and GTP, in the vacant site, could have biological implications in regulatory functions. Formation of this unique binding pocket in the G-triad could be used as a specific target in drug design.
- School of Physical and Mathematical Sciences, Nanyang Technological University, Singapore heddibrahim@ntu.edu.sg.
Organizational Affiliation: