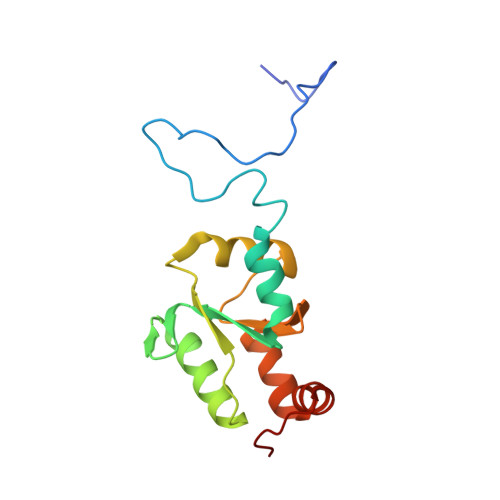
The lineage-specific, intrinsically disordered N-terminal extension of monothiol glutaredoxin 1 from trypanosomes contains a regulatory region.
Sturlese, M., Manta, B., Bertarello, A., Bonilla, M., Lelli, M., Zambelli, B., Grunberg, K., Mammi, S., Comini, M.A., Bellanda, M.(2018) Sci Rep 8: 13716-13716
- PubMed: 30209332
- DOI: https://doi.org/10.1038/s41598-018-31817-4
- Primary Citation of Related Structures:
2MXN - PubMed Abstract:
Glutaredoxins (Grx) are small proteins conserved throughout all the kingdoms of life that are engaged in a wide variety of biological processes and share a common thioredoxin-fold. Among them, class II Grx are redox-inactive proteins involved in iron-sulfur (FeS) metabolism. They contain a single thiol group in their active site and use low molecular mass thiols such as glutathione as ligand for binding FeS-clusters. In this study, we investigated molecular aspects of 1CGrx1 from the pathogenic parasite Trypanosoma brucei brucei, a mitochondrial class II Grx that fulfills an indispensable role in vivo. Mitochondrial 1CGrx1 from trypanosomes differs from orthologues in several features including the presence of a parasite-specific N-terminal extension (NTE) whose role has yet to be elucidated. Previously we have solved the structure of a truncated form of 1CGrx1 containing only the conserved glutaredoxin domain but lacking the NTE. Our aim here is to investigate the effect of the NTE on the conformation of the protein. We therefore solved the NMR structure of the full-length protein, which reveals subtle but significant differences with the structure of the NTE-less form. By means of different experimental approaches, the NTE proved to be intrinsically disordered and not involved in the non-redox dependent protein dimerization, as previously suggested. Interestingly, the portion comprising residues 65-76 of the NTE modulates the conformational dynamics of the glutathione-binding pocket, which may play a role in iron-sulfur cluster assembly and delivery. Furthermore, we disclosed that the class II-strictly conserved loop that precedes the active site is critical for stabilizing the protein structure. So far, this represents the first communication of a Grx containing an intrinsically disordered region that defines a new protein subgroup within class II Grx.
- Department of Chemical Sciences, University of Padova, via Marzolo 1, 35131, Padova, Italy.
Organizational Affiliation: