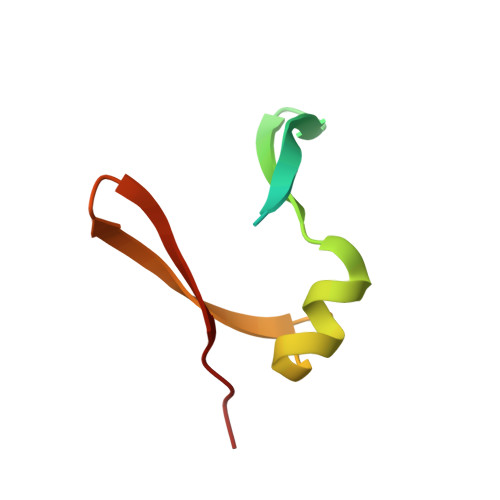
Escherichia coli antitoxin MazE as transcription factor: insights into MazE-DNA binding.
Zorzini, V., Buts, L., Schrank, E., Sterckx, Y.G., Respondek, M., Engelberg-Kulka, H., Loris, R., Zangger, K., van Nuland, N.A.(2015) Nucleic Acids Res 43: 1241-1256
- PubMed: 25564525 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gku1352
- Primary Citation Related Structures:
2MRN, 2MRU - PubMed Abstract:
Toxin-antitoxin (TA) modules are pairs of genes essential for bacterial regulation upon environmental stresses. The mazEF module encodes the MazF toxin and its cognate MazE antitoxin. The highly dynamic MazE possesses an N-terminal DNA binding domain through which it can negatively regulate its own promoter. Despite being one of the first TA systems studied, transcriptional regulation of Escherichia coli mazEF remains poorly understood. This paper presents the solution structure of C-terminal truncated E. coli MazE and a MazE-DNA model with a DNA palindrome sequence ∼ 10 bp upstream of the mazEF promoter. The work has led to a transcription regulator-DNA model, which has remained elusive thus far in the E. coli toxin-antitoxin family. Multiple complementary techniques including NMR, SAXS and ITC show that the long intrinsically disordered C-termini in MazE, required for MazF neutralization, does not affect the interactions between the antitoxin and its operator. Rather, the MazE C-terminus plays an important role in the MazF binding, which was found to increase the MazE affinity for the palindromic single site operator.
- Molecular Recognition Unit, Structural Biology Research Center, VIB, Pleinlaan 2, 1050 Brussels, Belgium Structural Biology Brussels, Vrije Universiteit Brussel, Pleinlaan 2, 1050 Brussels, Belgium.
Organizational Affiliation: