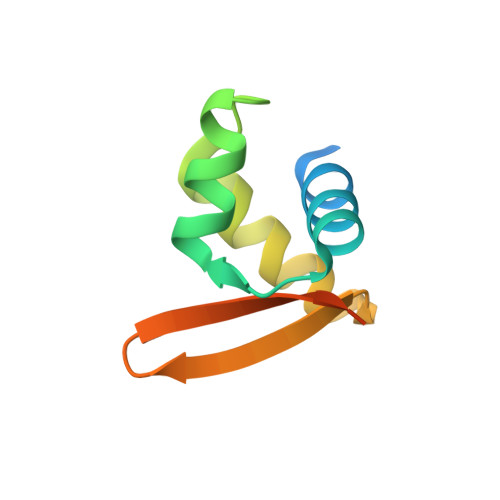
Solution Structure and DNA-binding Properties of the Winged Helix Domain of the Meiotic Recombination HOP2 Protein.
Moktan, H., Guiraldelli, M.F., Eyster, C.A., Zhao, W., Lee, C.Y., Mather, T., Camerini-Otero, R.D., Sung, P., Zhou, D.H., Pezza, R.J.(2014) J Biological Chem 289: 14682-14691
- PubMed: 24711446 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.548180
- Primary Citation Related Structures:
2MH2 - PubMed Abstract:
The HOP2 protein is required for efficient double-strand break repair which ensures the proper synapsis of homologous chromosomes and normal meiotic progression. We previously showed that in vitro HOP2 shows two distinctive activities: when it is incorporated into a HOP2-MND1 heterodimer, it stimulates DMC1 and RAD51 recombination activities, and the purified HOP2 alone is proficient in promoting strand invasion. The structural and biochemical basis of HOP2 action in recombination are poorly understood; therefore, they are the focus of this work. Herein, we present the solution structure of the amino-terminal portion of mouse HOP2, which contains a typical winged helix DNA-binding domain. Together with NMR spectral changes in the presence of double-stranded DNA, protein docking on DNA, and mutation analysis to identify the amino acids involved in DNA coordination, our results on the three-dimensional structure of HOP2 provide key information on the fundamental structural and biochemical requirements directing the interaction of HOP2 with DNA. These results, in combination with mutational experiments showing the role of a coiled-coil structural feature involved in HOP2 self-association, allow us to explain important aspects of the function of HOP2 in recombination.
- From the Department of Physics, Oklahoma State University, Stillwater, Oklahoma 74078.
Organizational Affiliation: