Structural basis and dynamics of multidrug recognition in a minimal bacterial multidrug resistance system
Habazettl, J., Allan, M., Jensen, P.R., Sass, H.J., Thompson, C.J., Grzesiek, S.(2014) Proc Natl Acad Sci U S A 111: E5498-E5507
- PubMed: 25489067 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1412070111
- Primary Citation Related Structures:
2MBZ, 2MC0 - PubMed Abstract:
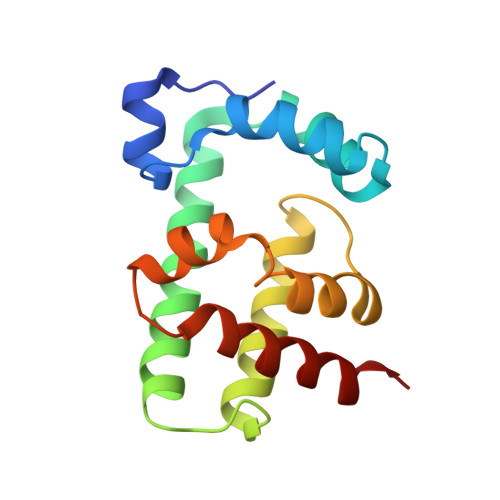
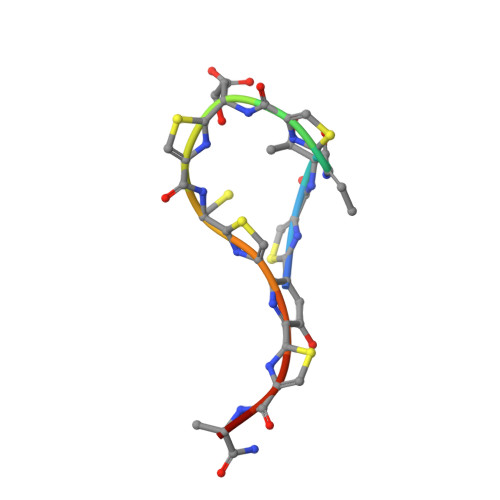
TipA is a transcriptional regulator found in diverse bacteria. It constitutes a minimal autoregulated multidrug resistance system against numerous thiopeptide antibiotics. Here we report the structures of its drug-binding domain TipAS in complexes with promothiocin A and nosiheptide, and a model of the thiostrepton complex. Drug binding induces a large transition from a partially unfolded to a globin-like structure. The structures rationalize the mechanism of promiscuous, yet specific, drug recognition: (i) a four-ring motif present in all known TipA-inducing antibiotics is recognized specifically by conserved TipAS amino acids; and (ii) the variable part of the antibiotic is accommodated within a flexible cleft that rigidifies upon drug binding. Remarkably, the identified four-ring motif is also the major interacting part of the antibiotic with the ribosome. Hence the TipA multidrug resistance mechanism is directed against the same chemical motif that inhibits protein synthesis. The observed identity of chemical motifs responsible for antibiotic function and resistance may be a general principle and could help to better define new leads for antibiotics.
- Focal Area Structural Biology and Biophysics, Biozentrum, University of Basel, CH-4056 Basel, Switzerland; and.
Organizational Affiliation: