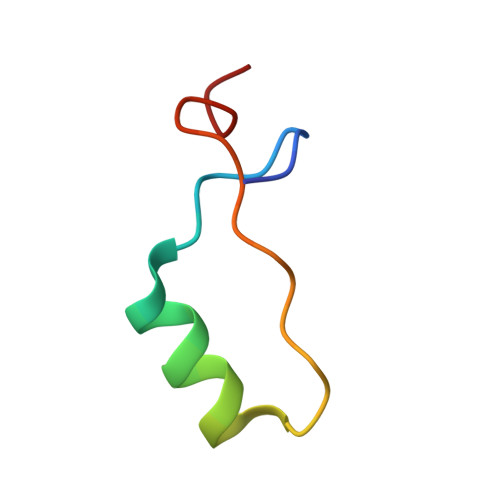
Solution Nuclear Magnetic Resonance Structure and Molecular Dynamics Simulations of a Murine 18.5 kDa Myelin Basic Protein Segment (S72-S107) in Association with Dodecylphosphocholine Micelles.
Ahmed, M.A., De Avila, M., Polverini, E., Bessonov, K., Bamm, V.V., Harauz, G.(2012) Biochemistry 51: 7475-7487
- PubMed: 22947219 Search on PubMed
- DOI: https://doi.org/10.1021/bi300998x
- Primary Citation Related Structures:
2LUG - PubMed Abstract:
The 18.5 kDa myelin basic protein (MBP), the most abundant splice isoform in adult mammalian myelin, is a multifunctional, intrinsically disordered protein involved in the development and compaction of the myelin sheath in the central nervous system. A highly conserved central segment comprises a membrane-anchoring amphipathic α-helix followed by a proline-rich segment that represents a ligand for SH3 domain-containing proteins. Here, we have determined using solution nuclear magnetic resonance spectroscopy the structure of a 36-residue peptide fragment of MBP (murine 18.5 kDa residues S72-S107, denoted the α2-peptide) comprising these two structural motifs, in association with dodecylphosphocholine (DPC) micelles. The structure was calculated using CS-ROSETTA (version 1.01) because the nuclear Overhauser effect restraints were insufficient for this protein. The experimental studies were complemented by molecular dynamics simulations of a corresponding 24-residue peptide fragment (murine 18.5 kDa residues E80-G103, denoted the MD-peptide), also in association with a DPC micelle in silico. The experimental and theoretical results agreed well with one another, despite the independence of the starting structures and analyses, both showing membrane association via the amphipathic α-helix, and a sharp bend in the vicinity of the Pro93 residue (murine 18.5 kDa sequence numbering). Overall, the conformations elucidated here show how the SH3 ligand is presented to the cytoplasm for interaction with SH3 domain-containing proteins such as Fyn and contribute to our understanding of myelin architecture at the molecular level.
- Department of Molecular and Cellular Biology, University of Guelph, 50 Stone Road East, Guelph, Ontario N1G 2W1, Canada.
Organizational Affiliation: