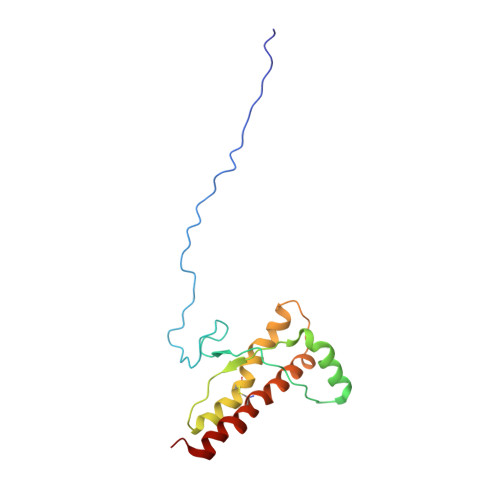
Toward the Molecular Basis of Inherited Prion Diseases: NMR Structure of the Human Prion Protein with V210I Mutation.
Biljan, I., Ilc, G., Giachin, G., Raspadori, A., Zhukov, I., Plavec, J., Legname, G.(2011) J Mol Biology 412: 660-673
- PubMed: 21839748 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.07.067
- Primary Citation Related Structures:
2LEJ - PubMed Abstract:
The development of transmissible spongiform encephalopathies (TSEs) is associated with the conversion of the cellular prion protein (PrP(C)) into a misfolded, pathogenic isoform (PrP(Sc)). Spontaneous generation of PrP(Sc) in inherited forms of disease is caused by mutations in gene coding for PrP (PRNP). In this work, we describe the NMR solution-state structure of the truncated recombinant human PrP (HuPrP) carrying the pathological V210I mutation linked to genetic Creutzfeldt-Jakob disease. The three-dimensional structure of V210I mutant consists of an unstructured N-terminal part (residues 90-124) and a well-defined C-terminal domain (residues 125-228). The C-terminal domain contains three α-helices (residues 144-156, 170-194 and 200-228) and a short antiparallel β-sheet (residues 129-130 and 162-163). Comparison with the structure of the wild-type HuPrP revealed that although two structures share similar global architecture, mutation introduces some local structural differences. The observed variations are mostly clustered in the α(2)-α(3) inter-helical interface and in the β(2)-α(2) loop region. Introduction of bulkier Ile at position 210 induces reorientations of several residues that are part of hydrophobic core, thus influencing α(2)-α(3) inter-helical interactions. Another important structural feature involves the alteration of conformation of the β(2)-α(2) loop region and the subsequent exposure of hydrophobic cluster to solvent, which facilitates intermolecular interactions involved in spontaneous generation of PrP(Sc). The NMR structure of V210I mutant offers new clues about the earliest events of the pathogenic conversion process that could be used for the development of antiprion drugs.
- Slovenian NMR Centre, National Institute of Chemistry, Hajdrihova 19, SI-1000 Ljubljana, Slovenia.
Organizational Affiliation: