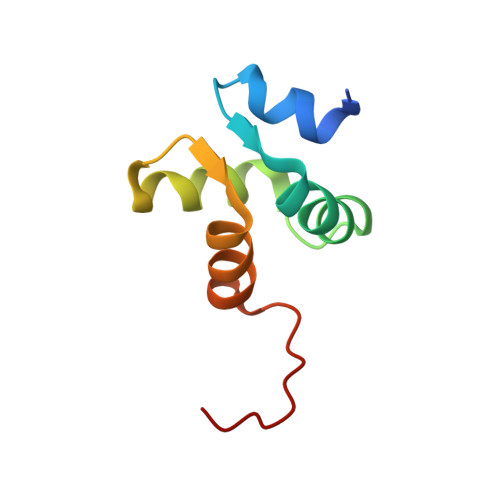
Solution structure of the N-terminal domain of the archaeal D-family DNA polymerase small subunit reveals evolutionary relationship to eukaryotic B-family polymerases
Yamasaki, K., Urushibata, Y., Yamasaki, T., Arisaka, F., Matsui, I.(2010) FEBS Lett 584: 3370-3375
- PubMed: 20598295
- DOI: https://doi.org/10.1016/j.febslet.2010.06.026
- Primary Citation Related Structures:
2KXE - PubMed Abstract:
Archaea-specific D-family DNA polymerase forms a heterotetramer consisting of two large polymerase subunits and two small exonuclease subunits. We analyzed the structure of the N-terminal 200 amino-acid regulatory region of the small subunit by NMR and revealed that the N-terminal approximately 70 amino-acid region is folded. The structure consists of a four-alpha-helix bundle including a short parallel beta-sheet, which is similar to the N-terminal regions of the B subunits of human DNA polymerases alpha and epsilon, establishing evolutionary relationships among these archaeal and eukaryotic polymerases. We observed monomer-dimer equilibrium of this domain, which may be related to holoenzyme architecture and/or functional regulation.
- Biomedical Research Institute, National Institute of Advanced Industrial Science and Technology (AIST), 1-1-1 Higashi, Tsukuba 305-8566, Japan. k-yamasaki@aist.go.jp
Organizational Affiliation: