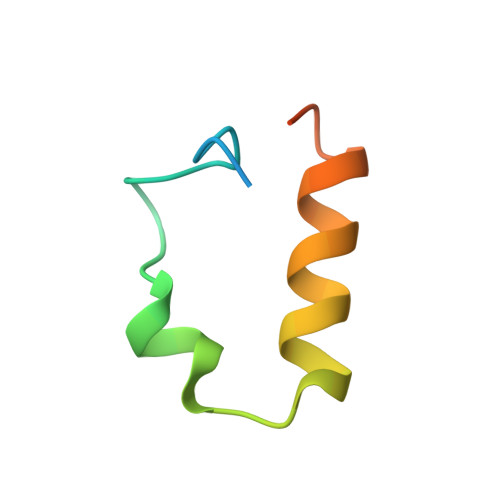
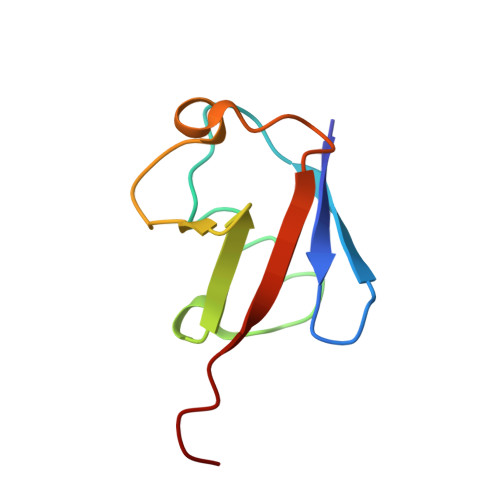
Structural analysis of the conserved ubiquitin-binding motifs (UBMs) of the translesion polymerase iota in complex with ubiquitin.
Burschowsky, D., Rudolf, F., Rabut, G., Herrmann, T., Matthias, P., Wider, G.(2011) J Biological Chem 286: 1364-1373
- PubMed: 20929865 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.135038
- Primary Citation Related Structures:
2KWU, 2KWV - PubMed Abstract:
Ubiquitin-binding domains (UBDs) provide specificity to the ubiquitin system, which is also involved in translesion synthesis (TLS) in eukaryotic cells. Upon DNA damage, the UBDs (UBM domains) of polymerase iota (Pol ι) interact with ubiquitinated proliferating cell nuclear antigen to regulate the interchange between processive DNA polymerases and TLS. We report a biophysical analysis and solution structures of the two conserved UBM domains located in the C-terminal tail of murine Pol ι in complex with ubiquitin. The 35-amino acid core folds into a helix-turn-helix motif, which belongs to a novel domain fold. Similar to other UBDs, UBMs bind to ubiquitin on the hydrophobic surface delineated by Leu-8, Ile-44, and Val-70, however, slightly shifted toward the C terminus. In addition, UBMs also use electrostatic interactions to stabilize binding. NMR and fluorescence spectroscopy measurements revealed that UBMs bind monoubiquitin, and Lys-63- but not Lys-48-linked chains. Importantly, these biophysical data are supported by functional studies. Indeed, yeast cells expressing ubiquitin mutants specifically defective for UBM binding are viable but sensitive to DNA damaging conditions that require TLS for repair.
- Institute of Molecular Biology and Biophysics, ETH Zurich, 8093 Zurich, Switzerland.
Organizational Affiliation: