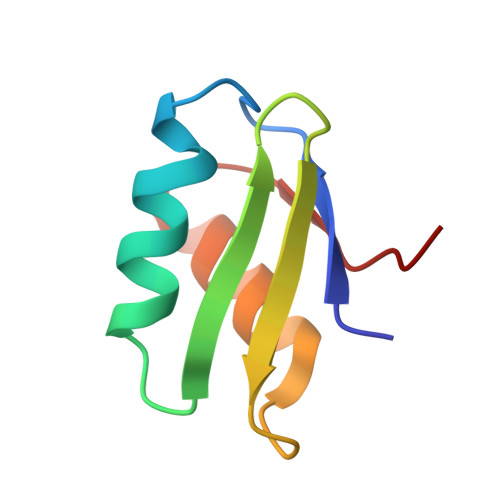
NmerA of Tn501 mercuric ion reductase: structural modulation of the pKa values of the metal binding cysteine thiols.
Ledwidge, R., Hong, B., Dotsch, V., Miller, S.M.(2010) Biochemistry 49: 8988-8998
- PubMed: 20828160
- DOI: https://doi.org/10.1021/bi100537f
- Primary Citation Related Structures:
2KT2, 2KT3 - PubMed Abstract:
To avoid nonspecific and/or undesirable binding and reactivity of metal ions with cellular components, organisms have evolved metal-specific systems for trafficking proteins. Although systems differ, those handling soft metal ions such as Hg(2+), Cu(+), Zn(2+), etc., all utilize heavy metal-associated (HMA) proteins and domains of ~70 amino acids with a conserved GMXCXXC motif in a βαββαβ structural fold. While the conserved cysteines define a common metal binding site in these proteins, other structural features must be utilized to create metal ion, protein partner, and contextual specificities. This paper presents initial structure-function studies of the N-terminal HMA domain (NmerA) of Tn501 mercuric ion reductase (MerA) aimed at identifying structural features critical to its role in facilitating efficient transfer of Hg(2+) to the MerA catalytic core for reductive detoxification. First, NMR solution structures of reduced and Hg(2+)-bound forms of NmerA are presented that allow definition and comparison of the structure of the metal binding loop in the two states. Structural differences between the two forms are compared with differences observed in three HMA domains with different metal ion and functional contexts. Second, analyses of the UV absorbance properties of wild-type, Cys11Ala, and Cys14Ala forms of NmerA are presented that provide assignments of the pK(a) values for the two cysteine thiols of the metal binding motif. Third, results from ¹³C NMR studies with wild-type and Y62F NmerA labeled with [β-¹³C]cysteine are presented that define a role for Tyr62 in modulating the pK(a) values of the cysteine thiols.
- Department of Pharmaceutical Chemistry, University of California-San Francisco, 600 16th Street, San Francisco, CA 94158-2517, USA.
Organizational Affiliation: