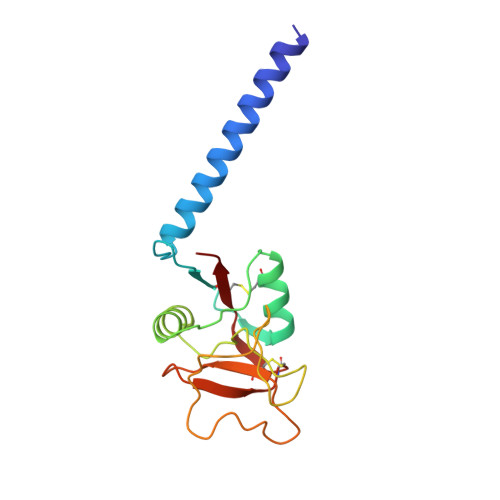
Structure of a selectin-like mutant of mannose-binding protein complexed with sialylated and sulfated Lewis(x) oligosaccharides.
Ng, K.K., Weis, W.I.(1997) Biochemistry 36: 979-988
- PubMed: 9033386 Search on PubMed
- DOI: https://doi.org/10.1021/bi962564e
- Primary Citation Related Structures:
1KMB, 2KMB, 3KMB, 4KMB - PubMed Abstract:
Rat serum mannose-binding protein in which residues 211-213 have been changed to the Lys-Lys-Lys sequence found in E-selectin binds HL-60 cells and the oligosaccharide 3'-NeuAc-Le(x). To understand how this mutant, designated K3, mimics the carbohydrate-binding properties of E-selectin, structures of K3 alone and in complexes with 3'-NeuAc-Le(x), 3'-sulfo-Le(x) and 4'-sulfo-Le(x) have been determined at 1.95-2.1 A resolution by X-ray crystallography. The region of K3 that interacts with bound oligosaccharides superimposes closely with the corresponding region of unliganded E-selectin. In each of the oligosaccharide-protein complexes, the 2- and 3-OH of Fuc coordinate Ca2+ and form a network of cooperative hydrogen bonds with amino acid side chains that also coordinate the Ca2+. Lys211 of the K3 mutant, which corresponds to Lys111 of E-selectin, interacts with each of the three bound ligands: the N zeta atom donates a hydrogen bond to the 4-OH of Gal in 3'-NeuAc-Le(x), forms a water-mediated hydrogen bond with the 4-OH of Gal in 3'-sulfo-Le(x), and forms a salt bridge with the sulfate group of 4'-sulfo-Le(x). Lys213 packs against an otherwise exposed aromatic residue and forms a water-mediated hydrogen bond with Lys211 which may help to position that residue for interactions with bound oligosaccharides. These structures are consistent with previous mutagenesis and chemical modification studies which demonstrate the importance of the Ca2+ ligands as well as Lys111 and Lys113 for carbohydrate binding in the selectins, and they provide a structural basis for understanding the selective recognition of negatively charged Le(x) derivatives by the selectins.
- Department of Structural Biology, Stanford University School of Medicine, California 94305, USA.
Organizational Affiliation: