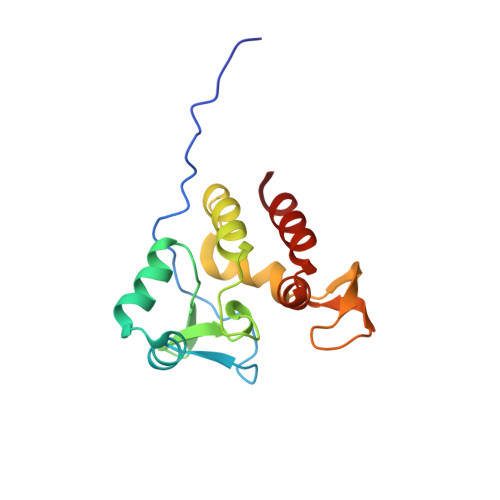
The Mycobacterium tuberculosis Ser/Thr Kinase Substrate Rv2175c Is a DNA-binding Protein Regulated by Phosphorylation.
Cohen-Gonsaud, M., Barthe, P., Canova, M.J., Stagier-Simon, C., Kremer, L., Roumestand, C., Molle, V.(2009) J Biological Chem 284: 19290-19300
- PubMed: 19457863 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.019653
- Primary Citation Related Structures:
2KFS - PubMed Abstract:
Recent efforts have underlined the role of serine/threonine protein kinases in growth, pathogenesis, and cell wall metabolism in Mycobacterium tuberculosis. Although most kinases have been investigated for their physiological roles, little information is available regarding how serine/threonine protein kinase-dependent phosphorylation regulates the activity of kinase substrates. Herein, we focused on M. tuberculosis Rv2175c, a protein of unknown function, conserved in actinomycetes, and recently identified as a substrate of the PknL kinase. We solved the solution structure of Rv2175c by multidimensional NMR and demonstrated that it possesses an original winged helix-turn-helix motif, indicative of a DNA-binding protein. The DNA-binding activity of Rv2175c was subsequently confirmed by fluorescence anisotropy, as well as in electrophoretic mobility shift assays. Mass spectrometry analyses using a combination of MALDI-TOF and LC-ESI/MS/MS identified Thr(9) as the unique phosphoacceptor. This was further supported by complete loss of PknL-dependent phosphorylation of an Rv2175c_T9A mutant. Importantly, the DNA-binding activity was completely abrogated in a Rv2175c_T9D mutant, designed to mimic constitutive phosphorylation, but not in a mutant lacking the first 13 residues. This implies that the function of the N-terminal extension is to provide a phosphoacceptor (Thr(9)), which, following phosphorylation, negatively regulates the Rv2175c DNA-binding activity. Interestingly, the N-terminal disordered extension, which bears the phosphoacceptor, was found to be restricted to members of the M. tuberculosis complex, thus suggesting the existence of an original mechanism that appears to be unique to the M. tuberculosis complex.
- CNRS UMR 5048, Centre de Biochimie Structurale, Montpellier, France. martin@cbs.cnrs.fr
Organizational Affiliation: