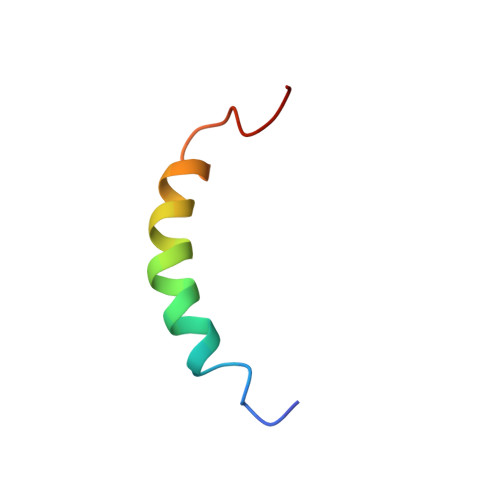
Three-dimensional structure of the two-peptide bacteriocin plantaricin JK.
Rogne, P., Haugen, C., Fimland, G., Nissen-Meyer, J., Kristiansen, P.E.(2009) Peptides 30: 1613-1621
- PubMed: 19538999
- DOI: https://doi.org/10.1016/j.peptides.2009.06.010
- Primary Citation of Related Structures:
2KEG, 2KEH, 2KHF, 2KHG - PubMed Abstract:
The three-dimensional structures of the two peptides, PlnJ and PlnK, that constitutes the two-peptide bacteriocin plantaricin JK have been solved in water/TFE and water/DPC-micellar solutions using nuclear magnetic resonance (NMR) spectroscopy. PlnJ, a 25 residue peptide, has an N-terminal amphiphilic alpha-helix between Trp-3 and Tyr-15. The 32 residues long PlnK forms a central amphiphilic alpha-helix between Gly-9 and Leu-24. Measurements of the effect on anti-microbial activity of single glycine replacements in PlnJ and PlnK show that Gly-13 and Gly-17 in both peptides are very sensitive, giving more than a 100-fold reduction in activity when large residues replace glycine. In variants where other glycine residues, Gly-20 in PlnJ and Gly-7, Gly-9, Gly-24 and Gly-25 in PlnK, were replaced, the activity was reduced less than 10-fold. It is proposed that the detrimental effect on activity when exchanging Gly-13 and Gly-17 in PlnJ and PlnK is a result of reduced ability of the two peptides to interact through the GxxxG-motifs constituting Gly-13 and Gly-17.
- Department of Molecular Biosciences, University of Oslo, Blindern, Oslo, Norway. p.a.rogne@imbv.uio.no
Organizational Affiliation: