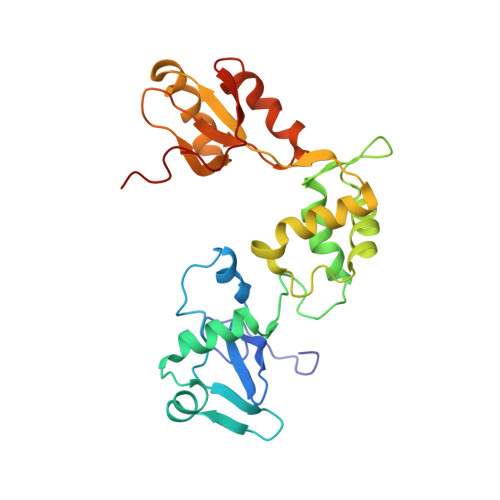
Structure, dynamics, and RNA interaction analysis of the human SBDS protein.
de Oliveira, J.F., Sforca, M.L., Blumenschein, T.M., Goldfeder, M.B., Guimaraes, B.G., Oliveira, C.C., Zanchin, N.I., Zeri, A.C.(2010) J Mol Biology 396: 1053-1069
- PubMed: 20053358 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.12.039
- Primary Citation Related Structures:
2KDO - PubMed Abstract:
Shwachman-Bodian-Diamond syndrome is an autosomal recessive genetic syndrome with pleiotropic phenotypes, including pancreatic deficiencies, bone marrow dysfunctions with increased risk of myelodysplasia or leukemia, and skeletal abnormalities. This syndrome has been associated with mutations in the SBDS gene, which encodes a conserved protein showing orthologs in Archaea and eukaryotes. The Shwachman-Bodian-Diamond syndrome pleiotropic phenotypes may be an indication of different cell type requirements for a fully functional SBDS protein. RNA-binding activity has been predicted for archaeal and yeast SBDS orthologs, with the latter also being implicated in ribosome biogenesis. However, full-length SBDS orthologs function in a species-specific manner, indicating that the knowledge obtained from model systems may be of limited use in understanding major unresolved issues regarding SBDS function, namely, the effect of mutations in human SBDS on its biochemical function and the specificity of RNA interaction. We determined the solution structure and backbone dynamics of the human SBDS protein and describe its RNA binding site using NMR spectroscopy. Similarly to the crystal structures of Archaea, the overall structure of human SBDS comprises three well-folded domains. However, significant conformational exchange was observed in NMR dynamics experiments for the flexible linker between the N-terminal domain and the central domain, and these experiments also reflect the relative motions of the domains. RNA titrations monitored by heteronuclear correlation experiments and chemical shift mapping analysis identified a classic RNA binding site at the N-terminal FYSH (fungal, Yhr087wp, Shwachman) domain that concentrates most of the mutations described for the human SBDS.
- Center for Structural Molecular Biology, Brazilian Synchrotron Light Laboratory, LNLS Rua Giuseppe Maximo Scolfaro 10000, PO Box 6192, CEP 13083-970 Campinas, SP, Brazil.
Organizational Affiliation: