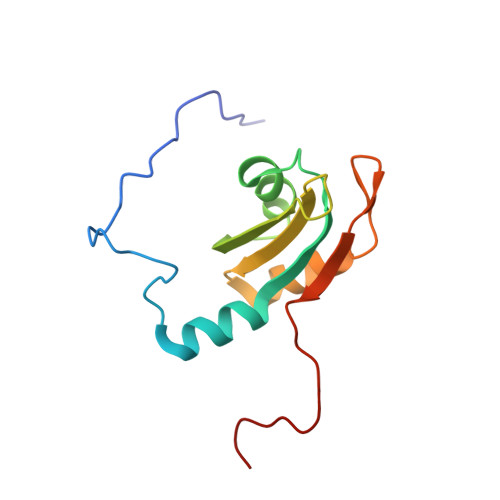
Structural basis for RNA recognition by a type II poly(A)-binding protein.
Song, J., McGivern, J.V., Nichols, K.W., Markley, J.L., Sheets, M.D.(2008) Proc Natl Acad Sci U S A 105: 15317-15322
- PubMed: 18824697 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0801274105
- Primary Citation Related Structures:
2JWN - PubMed Abstract:
We identified a functional domain (XlePABP2-TRP) of Xenopus laevis embryonic type II poly(A)-binding protein (XlePABP2). The NMR structure of XlePABP2-TRP revealed that the protein is a homodimer formed by the antiparallel association of beta-strands from the single RNA recognition motif (RRM) domain of each subunit. In each subunit of the homodimer, the canonical RNA recognition site is occluded by a polyproline motif. Upon poly(A) binding, XlePABP2-TRP undergoes a dimer-monomer transition that removes the polyproline motif from the RNA recognition site and allows it to be replaced by the adenosine nucleotides of poly(A). Our results provide high-resolution structural information concerning type II PABPs and an example of a single RRM domain protein that transitions from a homodimer to a monomer upon RNA binding. These findings advance our understanding of RRM domain regulation, poly(A) recognition, and are relevant to understanding how type II PABPs function in mRNA processing and human disease.
- Department of Biochemistry, Center for Eukaryotic Structural Genomics, University of Wisconsin, Madison, WI 53706, USA.
Organizational Affiliation: