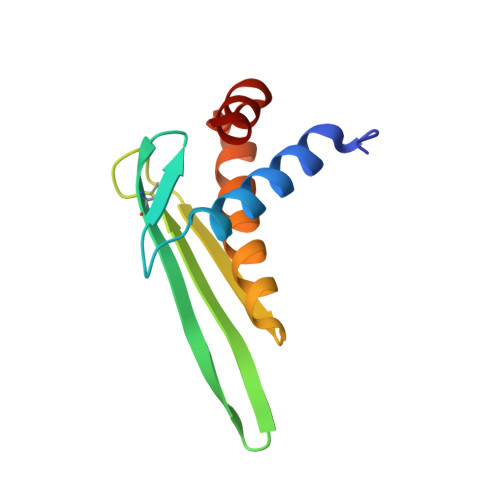
NMR structure of rALF-Pm3, an anti-lipopolysaccharide factor from shrimp: Model of the possible lipid A-binding site
Yang, Y., Boze, H., Chemardin, P., Padilla, A., Moulin, G., Tassanakajon, A., Pugniere, M., Roquet, F., Destoumieux-Garzon, D., Gueguen, Y., Bachere, E., Aumelas, A.(2009) Biopolymers 91: 207-220
- PubMed: 19107926
- DOI: https://doi.org/10.1002/bip.21119
- Primary Citation Related Structures:
2JOB - PubMed Abstract:
The anti-lipopolysaccharide factor ALF-Pm3 is a 98-residue protein identified in hemocytes from the black tiger shrimp Penaeus monodon. It was expressed in Pichia pastoris from the constitutive glyceraldehyde-3-phosphate dehydrogenase promoter as a folded and (15)N uniformly labeled rALF-Pm3 protein. Its 3D structure was established by NMR and consists of three alpha-helices packed against a four-stranded beta-sheet. The C(34)-C(55) disulfide bond was shown to be essential for the structure stability. By using surface plasmon resonance, we demonstrated that rALF-Pm3 binds to LPS, lipid A and to OM-174, a soluble analogue of lipid A. Biophysical studies of rALF-Pm3/LPS and rALF-Pm3/OM-174 complexes indicated rather high molecular sized aggregates, which prevented us to experimentally determine by NMR the binding mode of these lipids to rALF-Pm3. However, on the basis of striking structural similarities to the FhuA/LPS complex, we designed an original model of the possible lipid A-binding site of ALF-Pm3. Such a binding site, located on the ALF-Pm3 beta-sheet and involving seven charged residues, is well conserved in ALF-L from Limulus polyphemus and in ALF-T from Tachypleus tridentatus. In addition, our model is in agreement with experiments showing that beta-hairpin synthetic peptides corresponding to ALF-L beta-sheet bind to LPS. Delineating lipid A-binding site of ALFs will help go further in the de novo design of new antibacterial or LPS-neutralizing drugs.
- CNRS UMR5048, INSERM, U554, Université Montpellier 1 et 2, Centre de Biochimie Structurale, 29 rue de Navacelles, 34090 Montpellier, Cedex 9, France.
Organizational Affiliation: