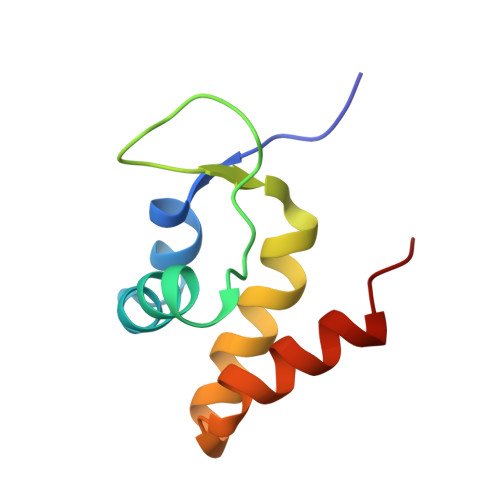
Structural basis for operator and antirepressor recognition by Myxococcus xanthus CarA repressor.
Navarro-Aviles, G., Jimenez, M.A., Perez-Marin, M.C., Gonzalez, C., Rico, M., Murillo, F.J., Elias-Arnanz, M., Padmanabhan, S.(2007) Mol Microbiol 63: 980-994
- PubMed: 17233828 Search on PubMed
- DOI: https://doi.org/10.1111/j.1365-2958.2006.05567.x
- Primary Citation Related Structures:
2JML - PubMed Abstract:
Blue light induces carotenogenesis in Myxococcus xanthus. The carB operon encodes all but one of the structural genes involved, and its expression is regulated by the CarA-CarS repressor-antirepressor pair. In the dark, CarA-operator binding represses carB. CarS, produced on illumination, interacts physically with CarA to dismantle the CarA-operator complex and activate carB. Both operator and CarS bind to the autonomously folded N-terminal domain of CarA, CarA(Nter), which in excess represses carB. Here, we report the NMR structure of CarA(Nter), and map residues that interact with operator and CarS by NMR chemical shift perturbations, and in vivo and in vitro analyses of site-directed mutants. We show CarA(Nter) adopts the winged-helix topology of MerR-family DNA-binding domains, and conserves the majority of the helix-turn-helix and wing contacts with DNA. Tellingly, helix alpha2 in CarA, a key element in operator DNA recognition, is also critical for interaction with CarS, implying that the CarA-CarS protein-protein and the CarA-operator protein-DNA interfaces overlap. Thus, binding of CarA to operator and to antirepressor are mutually exclusive, and CarA may discern structural features in the acidic CarS protein that resemble operator DNA. Repressor inactivation by occluding the DNA-binding region may be a recurrent mechanism of action for acidic antirepressors.
- Departamento de Genética y Microbiología, Facultad de Biología, Universidad de Murcia, Murcia, Spain.
Organizational Affiliation: