A Structural Explanation for the Binding of Endocytic Dileucine Motifs by the Ap2 Complex.
Kelly, B.T., Mccoy, A.J., Spaete, K., Miller, S.E., Evans, P.R., Hoening, S., Owen, D.J.(2008) Nature 456: 976
- PubMed: 19140243 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature07422
- Primary Citation Related Structures:
2JKR, 2JKT - PubMed Abstract:
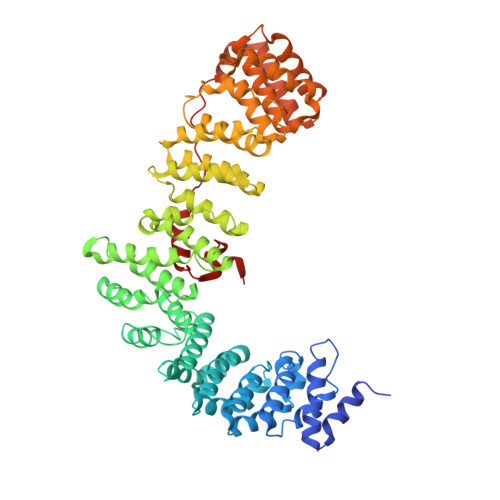
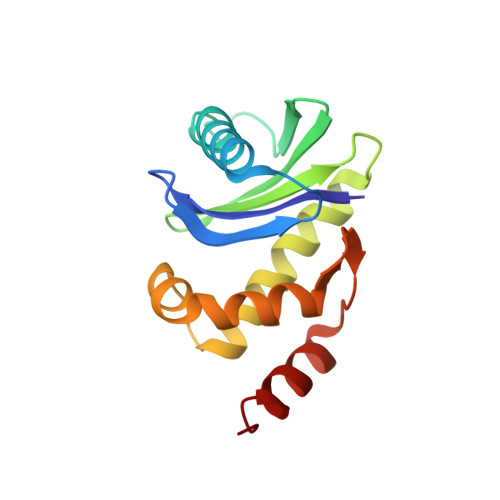
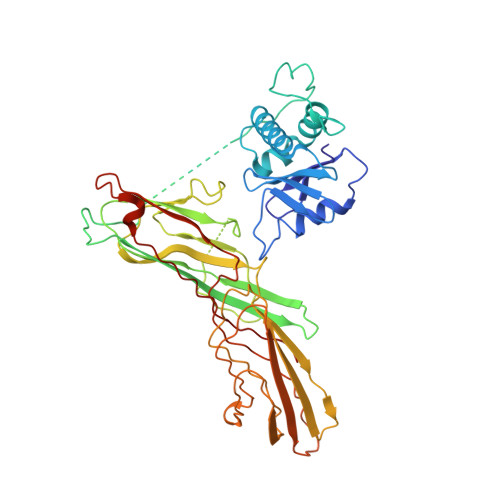
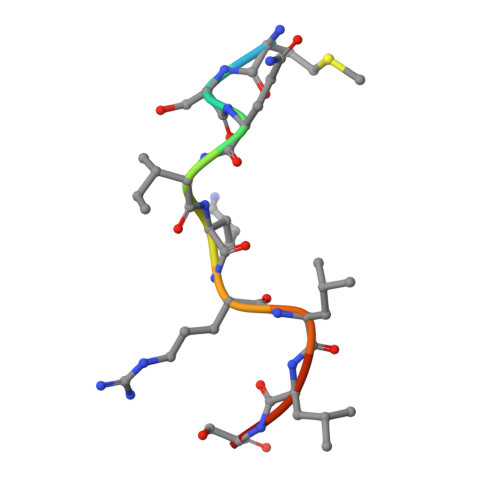
Most transmembrane proteins are selected as transport vesicle cargo through the recognition of short, linear amino acid motifs in their cytoplasmic portions by vesicle coat proteins. In the case of clathrin-coated vesicles (CCVs) the motifs are recognised by clathrin adaptors. The AP2 adaptor complex (subunits α,β2,μ2,σ2) recognises both major endocytic motifs: YxxΦ motifs and [DE]xxxL[LI] acidic dileucine motifs. Here we describe the binding of AP2 to the endocytic dileucine motif from CD4 . The major recognition events are the two leucine residues binding in hydrophobic pockets on σ2. The hydrophilic residue four residues upstream from the first leucine sits on a positively charged patch made from residues on σ2 and α subunits. Mutations in key residues inhibit the binding of AP2 to ‘acidic dileucine’ motifs displayed in liposomes containing PtdIns4,5P 2 , but do not affect binding to YxxΦ motifs via μ2. In the ‘inactive’ AP2 core structure , both motif binding sites are blocked by different parts of the β2 subunit. To allow a dileucine motif to bind, the β2 N-terminus is displaced and becomes disordered; however, in this structure the YxxΦ binding site on μ2 remains blocked.
- Cambridge Institute for Medical Research and Department of Clinical Biochemistry, University of Cambridge, Addenbrooke's Hospital, Hills Road, Cambridge. CB2 0XY, UK.
Organizational Affiliation: