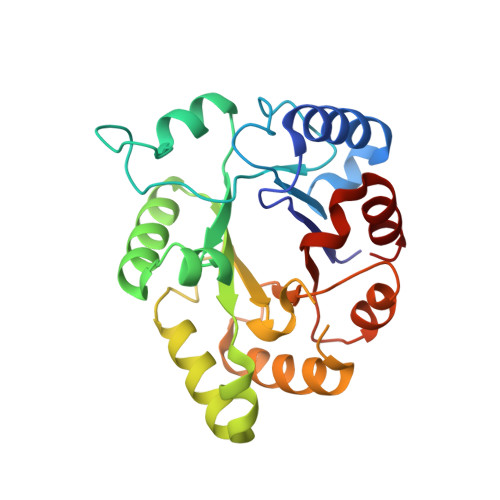
Kinetics and Structural Properties of Triosephosphate Isomerase from Helicobacter Pylori
Chu, C.-H., Lai, Y.-J., Sun, Y.-J.(2008) Proteins 71: 396
- PubMed: 17957775 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21709
- Primary Citation Related Structures:
2JGQ - PubMed Abstract:
Triosephosphate isomerase (TIM) catalyzes the interconversion between dihydroxyacetone phosphate and D-glyceraldehyde-3-phosphate in the glycolysis-gluconeogenesis metabolism pathway. The Helicobacter pylori TIM gene (HpTIM) was cloned, and HpTIM was expressed and purified. The enzymatic activity of HpTIM for the substrate GAP was determined (K(m) = 3.46 +/- 0.23 mM and k(cat) = 8.8 x 10(4) min(-1)). The crystal structure of HpTIM was determined by molecular replacement at 2.3 A resolution. The overall structure of HpTIM was (beta/alpha)beta(beta/alpha)(6), which resembles the common TIM barrel fold, (beta/alpha)(8); however, a helix is missing after the second beta-strand. The conformation of loop 6 and binding of phosphate ion suggest that the determined structure of HpTIM was in the "closed" state. A highly conserved Arg-Asp salt bridge in the "DX(D/N)G" motif of most TIMs is absent in HpTIM because the sequence of this motif is "(211)SVDG(214)." To determine the significance of this salt bridge to HpTIM, four mutants, including K183S, K183A, D213Q, and D213A, were constructed and characterized. The results suggest that this conserved salt bridge is not essential for the enzymatic activity of HpTIM; however, it might contribute to the conformational stability of HpTIM.
- Institute of Bioinformatics and Structural Biology, National Tsing Hua University, Hsinchu 30013, Taiwan, Republic of China.
Organizational Affiliation: