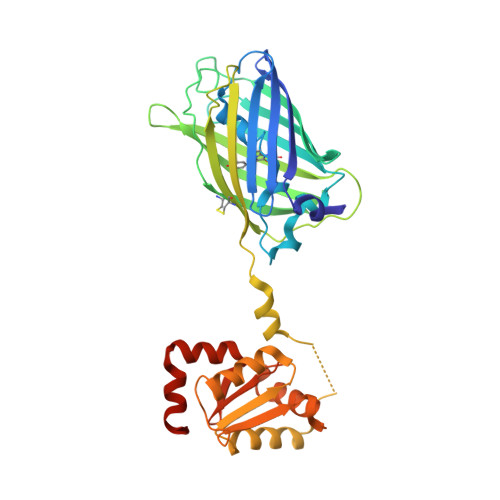
Structure of Glutaredoxin Grx1P C30S Mutant from Yeast.
Hakansson, K.O., Winther, J.R.(2007) Acta Crystallogr D Biol Crystallogr 63: 288
- PubMed: 17327665 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444906051675
- Primary Citation Related Structures:
2JAC, 2JAD - PubMed Abstract:
Glutathionylated glutaredoxin Grx1p C30S mutant from yeast has been crystallized in space group C222(1) and a fusion protein between redox-sensitive yellow fluorescent protein (rxYFP) and Grx1p C30S has been crystallized in space group P6(4). The structure of the latter was solved by molecular replacement using the known rxYFP structure as a search model. The structure of the Grx1p moiety was built and the structure was refined against 2.7 A synchrotron data to an R(free) of 25.7%. There are no specific contacts between the two domains, indicating that the observed enhanced exchange of reduction equivalents between them arises from diffusion or from an enhanced collision rate in solution. The Grx1p structure thus obtained was subsequently used to solve the structure of the orthorhombic crystal, which could be refined against 2.0 A data to an R(free) of 24.3%. The structure of the glutathione-bound protein and the glutaredoxin domain in the fusion protein are similar. The covalent disulfide bond between the glutathione and protein is broken upon exposure to synchrotron radiation. The structure and the glutathione-binding mode are described and compared with existing crystallographic and nuclear magnetic resonance (NMR) structures of related glutaredoxins. Conserved residues are clustered on one side of the active site.
- Department of Molecular Biology, August Krogh Building, University of Copenhagen, Universitetsparken 13, DK-2100 Copenhagen Ø, Denmark. kohakansson@aki.ku.dk
Organizational Affiliation: