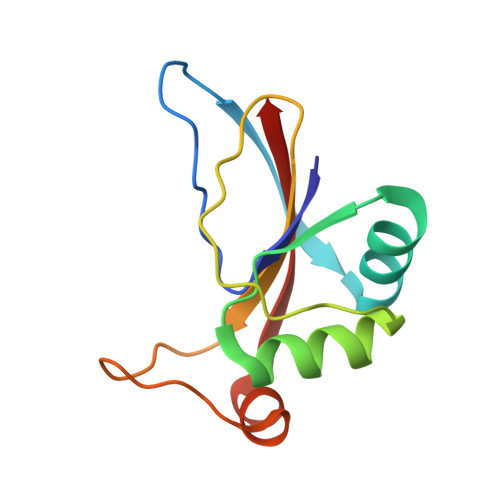
Crystal Structure of Afv3-109, a Highly Conserved Protein from Crenarchaeal Viruses.
Keller, J., Leulliot, N., Cambillau, C., Campanacci, V., Porciero, S., Prangishvili, D., Forterre, P., Cortez, D., Quevillon-Cheruel, S., Van Tilbeurgh, H.(2007) Virol J 4: 12
- PubMed: 17241456
- DOI: https://doi.org/10.1186/1743-422X-4-12
- Primary Citation Related Structures:
2J6B, 2J6C - PubMed Abstract:
The extraordinary morphologies of viruses infecting hyperthermophilic archaea clearly distinguish them from bacterial and eukaryotic viruses. Moreover, their genomes code for proteins that to a large extend have no related sequences in the extent databases. However, a small pool of genes is shared by overlapping subsets of these viruses, and the most conserved gene, exemplified by the ORF109 of the Acidianus Filamentous Virus 3, AFV3, is present on genomes of members of three viral familes, the Lipothrixviridae, Rudiviridae, and "Bicaudaviridae", as well as of the unclassified Sulfolobus Turreted Icosahedral Virus, STIV. We present here the crystal structure of the protein (Mr = 13.1 kD, 109 residues) encoded by the AFV3 ORF 109 in two different crystal forms at 1.5 and 1.3 A resolution. The structure of AFV3-109 is a five stranded beta-sheet with loops on one side and three helices on the other. It forms a dimer adopting the shape of a cradle that encompasses the best conserved regions of the sequence. No protein with a related fold could be identified except for the ortholog from STIV1, whose structure was deposited at the Protein Data Bank. We could clearly identify a well bound glycerol inside the cradle, contacting exclusively totally conserved residues. This interaction was confirmed in solution by fluorescence titration. Although the function of AFV3-109 cannot be deduced directly from its structure, structural homology with the STIV1 protein, and the size and charge distribution of the cavity suggested it could interact with nucleic acids. Fluorescence quenching titrations also showed that AFV3-109 interacts with dsDNA. Genomic sequence analysis revealed bacterial homologs of AFV3-109 as a part of a putative previously unidentified prophage sequences in some Firmicutes.
- Institut de Biochimie et de Biophysique Moléculaire et Cellulaire, CNRS-UMR 8619, Université Paris 11, IFR115, Bâtiment 430, 91405 Orsay, France. Jenny.Keller@u-psud.fr
Organizational Affiliation: