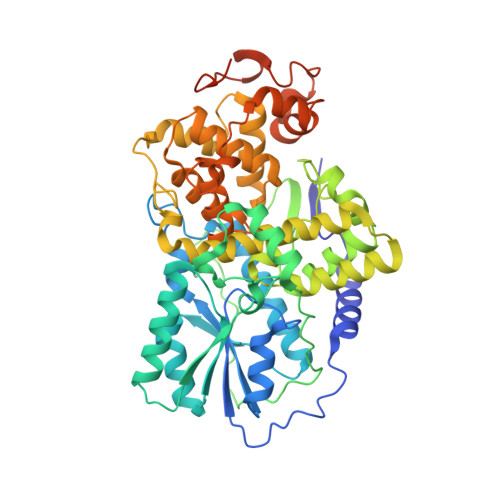
Cryptochrome 3 from Arabidopsis Thaliana: Structural and Functional Analysis of its Complex with a Folate Light Antenna
Klar, T., Pokorny, R., Moldt, J., Batschauer, A., Essen, L.-O.(2007) J Mol Biology 366: 954
- PubMed: 17188299 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.11.066
- Primary Citation Related Structures:
2J4D - PubMed Abstract:
Cryptochromes are almost ubiquitous blue-light receptors and act in several species as central components of the circadian clock. Despite being evolutionary and structurally related with DNA photolyases, a class of light-driven DNA-repair enzymes, and having similar cofactor compositions, cryptochromes lack DNA-repair activity. Cryptochrome 3 from the plant Arabidopsis thaliana belongs to the DASH-type subfamily. Its crystal structure determined at 1.9 Angstroms resolution shows cryptochrome 3 in a dimeric state with the antenna cofactor 5,10-methenyltetrahydrofolate (MTHF) bound in a distance of 15.2 Angstroms to the U-shaped FAD chromophore. Spectroscopic studies on a mutant where a residue crucial for MTHF-binding, E149, was replaced by site-directed mutagenesis demonstrate that MTHF acts in cryptochrome 3 as a functional antenna for the photoreduction of FAD.
- Philipps-Universität Marburg, Department of Chemistry, Hans-Meerwein-Strasse, D-35032 Marburg, Germany.
Organizational Affiliation: