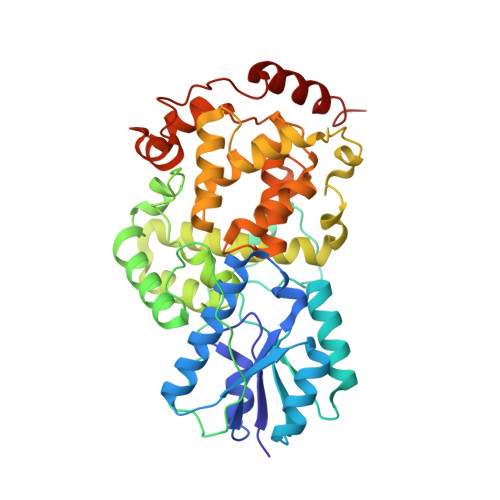
Natural and Non-Natural Antenna Chromophores in the DNA Photolyase from Thermus Thermophilus
Klar, T., Kaiser, G., Hennecke, U., Carell, T., Batschauer, A., Essen, L.-O.(2006) Chembiochem 7: 1798
- PubMed: 17051659 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.200600206
- Primary Citation Related Structures:
2J07, 2J08, 2J09 - PubMed Abstract:
X-ray crystallographic and functional analysis of the class I DNA photolyase from Thermus thermophilus revealed the binding of flavin mononucleotide (FMN) as an antenna chromophore. The binding mode of FMN closely coincides with the binding of a deazaflavin-like chromophore in the related class I DNA photolyase from Anacystis nidulans. Compared to the R46E mutant, which lacks a conserved arginine in the binding site for the antenna chromophore, the FMN-comprising holophotolyase exhibits an eightfold higher activity at 450 nm. The facile incorporation of the flavin cofactors 8-hydroxy-deazariboflavin and 8-iodo-8-demethyl-riboflavin into the binding site for the antenna chromophore paves the way for wavelength-tuning of the activity spectra of DNA photolyases by using synthetic flavins.
- Philipps-Universität Marburg, Fachbereich Chemie, Hans-Meerwein-Strasse, 35032 Marburg, Germany.
Organizational Affiliation: