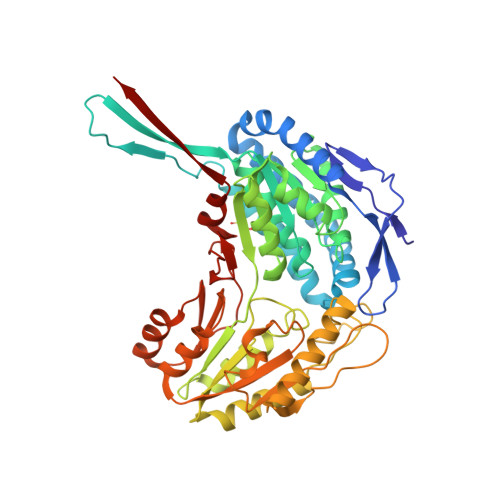
Crystal Structure of Lactaldehyde Dehydrogenase from Escherichia coli and Inferences Regarding Substrate and Cofactor Specificity.
Di Costanzo, L., Gomez, G.A., Christianson, D.W.(2007) J Mol Biology 366: 481-493
- PubMed: 17173928 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2006.11.023
- Primary Citation Related Structures:
2HG2, 2ILU, 2IMP - PubMed Abstract:
Aldehyde dehydrogenases catalyze the oxidation of aldehyde substrates to the corresponding carboxylic acids. Lactaldehyde dehydrogenase from Escherichia coli (aldA gene product, P25553) is an NAD(+)-dependent enzyme implicated in the metabolism of l-fucose and l-rhamnose. During the heterologous expression and purification of taxadiene synthase from the Pacific yew, lactaldehyde dehydrogenase from E. coli was identified as a minor (
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, PA 19104-6323, USA.
Organizational Affiliation: