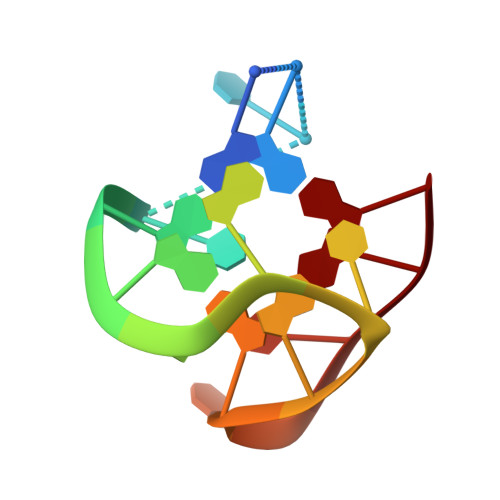
A new modified thrombin binding aptamer containing a 5'-5' inversion of polarity site.
Martino, L., Virno, A., Randazzo, A., Virgilio, A., Esposito, V., Giancola, C., Bucci, M., Cirino, G., Mayol, L.(2006) Nucleic Acids Res 34: 6653-6662
- PubMed: 17145716
- DOI: https://doi.org/10.1093/nar/gkl915
- Primary Citation Related Structures:
2IDN - PubMed Abstract:
The solution structure of a new modified thrombin binding aptamer (TBA) containing a 5'-5' inversion of polarity site, namely d(3'GGT5'-5'TGGTGTGGTTGG3'), is reported. NMR and CD spectroscopy, as well as molecular dynamic and mechanic calculations, have been used to characterize the 3D structure. The modified oligonucleotide is characterized by a chair-like structure consisting of two G-tetrads connected by three edge-wise TT, TGT and TT loops. d(3'GGT5'-5'TGGTGTGGTTGG3') is characterized by an unusual folding, being three strands parallel to each other and only one strand oriented in opposite manner. This led to an anti-anti-anti-syn and syn-syn-syn-anti arrangement of the Gs in the two tetrads. The thermal stability of the modified oligonucleotide is 4 degrees C higher than the corresponding unmodified TBA. d(3'GGT5'-5'TGGTGTGGTTGG3') continues to display an anticoagulant activity, even if decreased with respect to the TBA.
- Dipartimento di Chimica, Università degli Studi di Napoli Federico II, via Cintia, I-80126, Napoli, Italy.
Organizational Affiliation: