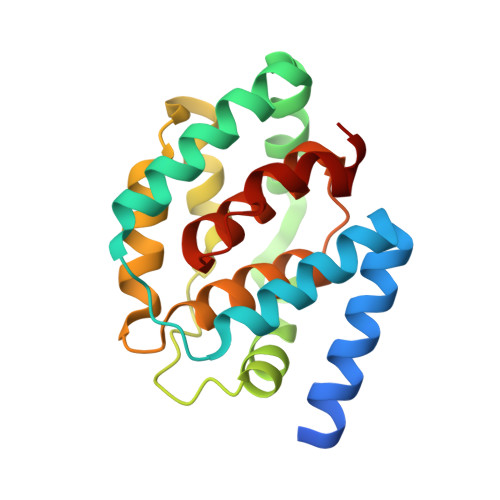
A mechanism for toxin insertion into membranes is suggested by the crystal structure of the channel-forming domain of colicin E1
Elkins, P., Bunker, A., Cramer, W.A., Stauffacher, C.V.(1997) Structure 5: 443-458
- PubMed: 9083117
- DOI: https://doi.org/10.1016/s0969-2126(97)00200-1
- Primary Citation Related Structures:
2I88 - PubMed Abstract:
Channel-forming colicins, including colicin E1, are a sub-family of bacteriocins. The toxic action of colicin E1 is derived from its ability to form a voltage-gated channel, which causes depolarization of the cytoplasmic membrane of sensitive Escherichia coli cells. In this process, the toxin-like colicin E1 molecule must undergo a substantial structural transition from a soluble state, in which it binds the target cell, to a membrane-bound state. Details of the structural changes that accompany this conversion may be directly applicable to other channel-forming toxins, as well as to the mechanism by which proteins insert into or cross membranes. The structure of the 190-residue channel-forming domain of colicin E1 in its soluble form has been solved at 2.5 A resolution. This structure contains 10alpha helices arranged in three layers (A-C) with a central hydrophobic helical hairpin in layer B, which is proposed to anchor the membrane-bound form in the bilayer. The extended N-terminal helix I provides a connection to the rest of the colicin E1 molecule, and the loop I-II may act as a hinge for re-orientation of the domain for membrane binding. A set of conserved positively charged residues on layer C may provide the docking surface on the molecule for membrane attachment. A large internal cavity between layers B and C may allow these layers to disengage, suggesting a mechanism for unfolding the molecule on the membrane that involves the perturbation of the interhelical hydrophobic interactions in layer C. On the basis of the structure of the colicin E1 channel-forming domain, its comparison with the structure of the colicin A domain and the known requirement for initial electrostatic and subsequent hydrophobic interactions, molecular details of the docking, unfolding and insertion of the channel-forming domain into the membrane are proposed. The model for docking and initial interaction with the membrane positions the hydrophobic hairpin 'anchor' approximately parallel to the membrane surface. Hydrophobic interactions in the docking layer may then be displaced by interactions with the membrane, spreading the helices on the surface and exposing the hydrophobic hairpin for insertion into the membrane.
- Protein Engineering, Department Genentech, Inc. 460 Pt. San Bruno Blvd, South San Francisco, CA 94080, USA.
Organizational Affiliation: