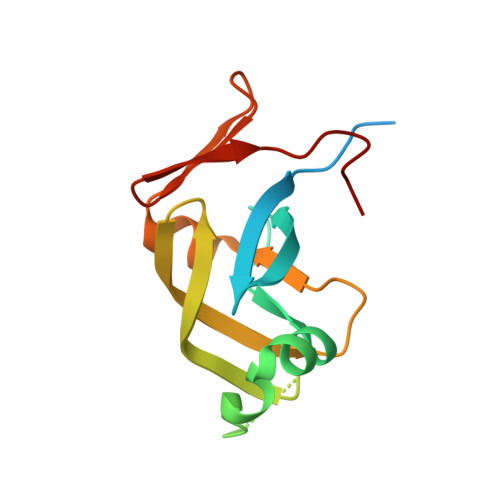
Ddi1, a eukaryotic protein with the retroviral protease fold.
Sirkis, R., Gerst, J.E., Fass, D.(2006) J Mol Biology 364: 376-387
- PubMed: 17010377
- DOI: https://doi.org/10.1016/j.jmb.2006.08.086
- Primary Citation Related Structures:
2I1A - PubMed Abstract:
Retroviral aspartyl proteases are homodimeric, whereas eukaryotic aspartyl proteases tend to be large, monomeric enzymes with 2-fold internal symmetry. It has been proposed that contemporary monomeric aspartyl proteases evolved by gene duplication and fusion from a primordial homodimeric enzyme. Recent sequence analyses have suggested that such "fossil" dimeric aspartyl proteases are still encoded in the eukaryotic genome. We present evidence for retention of a dimeric aspartyl protease in eukaryotes. The X-ray crystal structure of a domain of the Saccharomyces cerevisiae protein Ddi1 shows that it is a dimer with a fold similar to that of the retroviral proteases. Furthermore, the double Asp-Thr-Gly-Ala amino acid sequence motif at the active site of HIV protease is found with identical geometry in the Ddi1 structure. However, the putative substrate binding groove is wider in Ddi1 than in the retroviral proteases, suggesting that Ddi1 accommodates bulkier substrates. Ddi1 belongs to a family of proteins known as the ubiquitin receptors, which have in common the ability to bind ubiquitinated substrates and the proteasome. Ubiquitin receptors contain an amino-terminal ubiquitin-like (UBL) domain and a carboxy-terminal ubiquitin-associated (UBA) domain, but Ddi1 is the only representative in which the UBL and UBA domains flank an aspartyl protease-like domain. The remarkable structural similarity between the central domain of Ddi1 and the retroviral proteases, in the global fold and in active-site detail, suggests that Ddi1 functions proteolytically during regulated protein turnover in the cell.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: