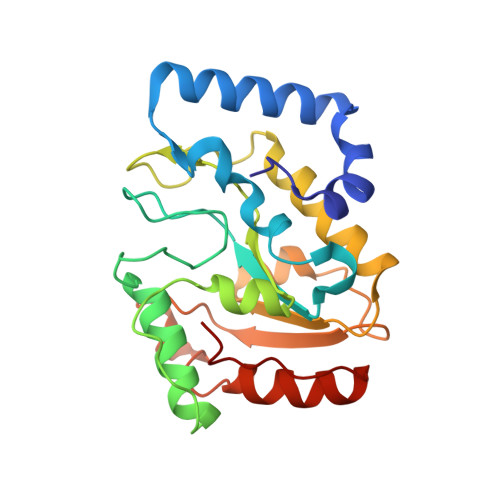
Mimicking damaged DNA with a small molecule inhibitor of human UNG2.
Krosky, D.J., Bianchet, M.A., Seiple, L., Chung, S., Amzel, L.M., Stivers, J.T.(2006) Nucleic Acids Res 34: 5872-5879
- PubMed: 17062624 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkl747
- Primary Citation Related Structures:
2HXM - PubMed Abstract:
Human nuclear uracil DNA glycosylase (UNG2) is a cellular DNA repair enzyme that is essential for a number of diverse biological phenomena ranging from antibody diversification to B-cell lymphomas and type-1 human immunodeficiency virus infectivity. During each of these processes, UNG2 recognizes uracilated DNA and excises the uracil base by flipping it into the enzyme active site. We have taken advantage of the extrahelical uracil recognition mechanism to build large small-molecule libraries in which uracil is tethered via flexible alkane linkers to a collection of secondary binding elements. This high-throughput synthesis and screening approach produced two novel uracil-tethered inhibitors of UNG2, the best of which was crystallized with the enzyme. Remarkably, this inhibitor mimics the crucial hydrogen bonding and electrostatic interactions previously observed in UNG2 complexes with damaged uracilated DNA. Thus, the environment of the binding site selects for library ligands that share these DNA features. This is a general approach to rapid discovery of inhibitors of enzymes that recognize extrahelical damaged bases.
- Department of Pharmacology and Molecular Sciences, 725 North Wolfe Street, Baltimore, MD 21205, USA.
Organizational Affiliation: