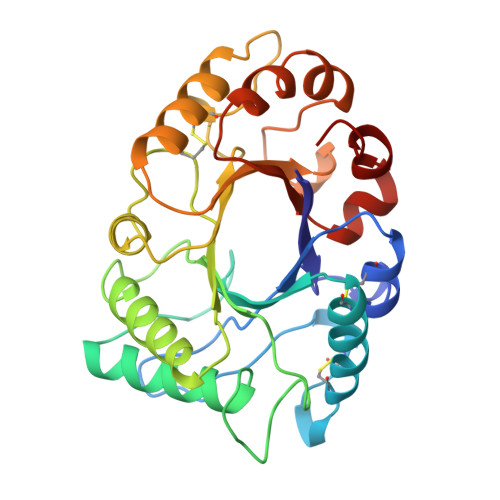
The 1.8 A resolution structure of hevamine, a plant chitinase/lysozyme, and analysis of the conserved sequence and structure motifs of glycosyl hydrolase family 18.
Terwisscha van Scheltinga, A.C., Hennig, M., Dijkstra, B.W.(1996) J Mol Biology 262: 243-257
- PubMed: 8831791
- DOI: https://doi.org/10.1006/jmbi.1996.0510
- Primary Citation Related Structures:
2HVM - PubMed Abstract:
The three-dimensional structure of hevamine, a plant enzyme with chitinase and lysozyme activity, has been refined at 1.8 A resolution to an R-factor of 14.9% and a free R-factor of 19.6%. The final model consists of all 273 amino acid residues and 206 ordered water molecules. Two non-proline cis-peptides were identified, involving Phe32 and Trp255, both of which are implicated in substrate binding. Other glycosyl hydrolase family 18 proteins with known three-dimensional structure are bacterial chitinase A, endo-beta-N-acetylglucosaminidase F1, endo-beta-N-acetylglucosaminidase H, and the two plant proteins concanavalin B and narbonin, which have no known enzymatic activity. All these structures contain a (beta alpha)8 barrel fold, with the two family 18 consensus regions roughly corresponding to the third and fourth barrel strands. This confirms the grouping of these proteins into family 18, which was only based on weak and local sequence similarity. The substrate specificity of the enzymes is determined by the loops following the barrel strands that form the substrate binding site. All enzymes have an aspartic acid and a glutamic acid residue in positions identical with Asp 125 and the catalytic Glu127 of hevamine. The lack of chitinase activity of concanavalin B and narbonin can be explained by the absence of one of these carboxylate groups, and by differences in the loops that form the substrate-binding cleft in hevamine.
- BIOSON Research Institute and Laboratory of Biophysical Chemistry, University of Groningen, The Netherlands.
Organizational Affiliation: