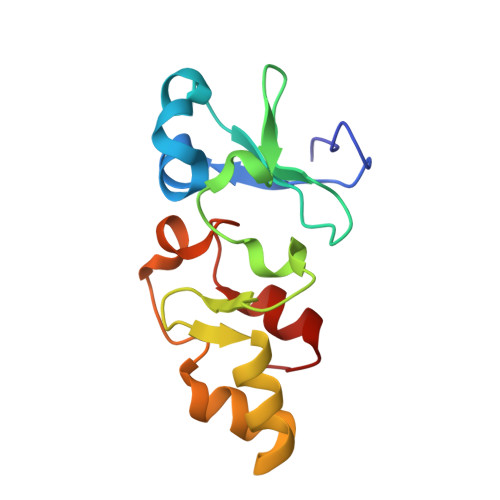
Characterization and structure of a Zn2+ and [2Fe-2S]-containing copper chaperone from Archaeoglobus fulgidus.
Sazinsky, M.H., LeMoine, B., Orofino, M., Davydov, R., Bencze, K.Z., Stemmler, T.L., Hoffman, B.M., Arguello, J.M., Rosenzweig, A.C.(2007) J Biological Chem 282: 25950-25959
- PubMed: 17609202 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M703311200
- Primary Citation Related Structures:
2HU9 - PubMed Abstract:
Bacterial CopZ proteins deliver copper to P1B-type Cu+-ATPases that are homologous to the human Wilson and Menkes disease proteins. The genome of the hyperthermophile Archaeoglobus fulgidus encodes a putative CopZ copper chaperone that contains an unusual cysteine-rich N-terminal domain of 130 amino acids in addition to a C-terminal copper binding domain with a conserved CXXC motif. The N-terminal domain (CopZ-NT) is homologous to proteins found only in extremophiles and is the only such protein that is fused to a copper chaperone. Surprisingly, optical, electron paramagnetic resonance, and x-ray absorption spectroscopic data indicate the presence of a [2Fe-2S] cluster in CopZ-NT. The intact CopZ protein binds two copper ions, one in each domain. The 1.8 A resolution crystal structure of CopZ-NT reveals that the [2Fe-2S] cluster is housed within a novel fold and that the protein also binds a zinc ion at a four-cysteine site. CopZ can deliver Cu+ to the A. fulgidus CopA N-terminal metal binding domain and is capable of reducing Cu2+ to Cu+. This unique fusion of a redox-active domain with a CXXC-containing copper chaperone domain is relevant to the evolution of copper homeostatic mechanisms and suggests new models for copper trafficking.
- Department of Biochemistry, Molecular Biology, and Cell Biology, Northwestern University, Evanston, Illinois 60208, USA.
Organizational Affiliation: