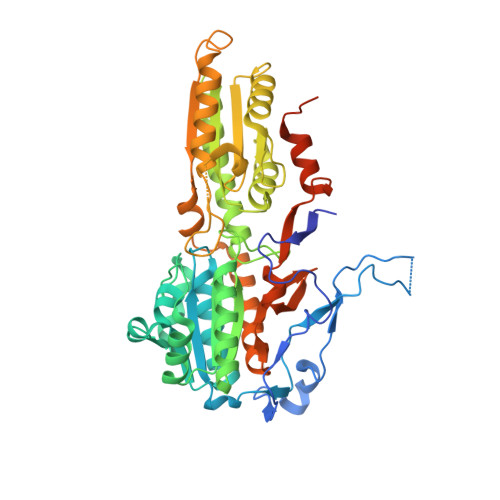
The First Crystal Structure of Phosphofructokinase from a Eukaryote: Trypanosoma brucei.
Martinez-Oyanedel, J., McNae, I.W., Nowicki, M.W., Keillor, J.W., Michels, P.A., Fothergill-Gilmore, L.A., Walkinshaw, M.D.(2007) J Mol Biology 366: 1185-1198
- PubMed: 17207816 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.10.019
- Primary Citation Related Structures:
2HIG - PubMed Abstract:
The crystal structure of the ATP-dependent phosphofructokinase (PFK) from Trypanosoma brucei provides the first detailed description of a eukaryotic PFK, and enables comparisons to be made with the crystal structures of bacterial ATP-dependent and PPi-dependent PFKs. The structure reveals that two insertions (the 17-20 and 329-348 loops) that are characteristic of trypanosomatid PFKs, but absent from bacterial and mammalian ATP-dependent PFKs, are located within and adjacent to the active site, and are in positions to play important roles in the enzyme's mechanism. The 90 residue N-terminal extension forms a novel domain that includes an "embracing arm" across the subunit boundary to the symmetry-related subunit in the tetrameric enzyme. Comparisons with the PPi-dependent PFK from Borrelia burgdorferi show that several features thought to be characteristic of PPi-dependent PFKs are present in the trypanosome ATP-dependent PFK. These two enzymes are generally more similar to each other than to the bacterial or mammalian ATP-dependent PFKs. However, there are critical differences at the active site of PPi-dependent PFKs that are sufficient to prevent the binding of ATP. This crystal structure of a eukaryotic PFK has enabled us to propose a detailed model of human muscle PFK that shows active site and other differences that offer opportunities for structure-based drug discovery for the treatment of sleeping sickness and other diseases caused by the trypanosomatid family of protozoan parasites.
- Structural Biochemistry Group, Institute of Structural and Molecular Biology, University of Edinburgh, King's Buildings, Edinburgh EH9 3JR, Scotland.
Organizational Affiliation: