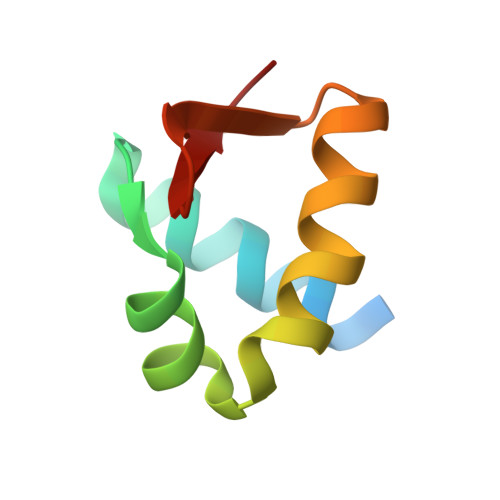
Structure-based secondary structure-independent approach to design protein ligands: Application to the design of Kv1.2 potassium channel blockers.
Magis, C., Gasparini, D., Lecoq, A., Le Du, M.H., Stura, E., Charbonnier, J.B., Mourier, G., Boulain, J.C., Pardo, L., Caruana, A., Joly, A., Lefranc, M., Masella, M., Menez, A., Cuniasse, P.(2006) J Am Chem Soc 128: 16190-16205
- PubMed: 17165772 Search on PubMed
- DOI: https://doi.org/10.1021/ja0646491
- Primary Citation Related Structures:
2HEO - PubMed Abstract:
We have developed a structure-based approach to the design of protein ligands. This approach is based on the transfer of a functional binding motif of amino acids, often referred as to the "hot spot", on a host protein able to reproduce the functional topology of these residues. The scaffolds were identified by a systematic in silico search in the Protein Data Bank for proteins possessing a group of residues in a topology similar to that adopted by the functional motif in a reference ligand of known 3D structure. In contrast to previously reported studies, this search is independent of the particular secondary structure supporting the functional motif. To take into account the global properties of the host protein, two additional criteria were taken into account in the selection process: (1) Only those scaffolds sterically compatible with the positioning of the functional motif as observed in a reference complex model were retained. (2) Host proteins displaying electrostatic potentials, in the region of the transferred functional motif, similar to that of the reference ligand were selected. This approach was applied to the development of protein ligands of the Kv1.2 channel using BgK, a small protein isolated from the sea anemone Bunodosoma granulifera, as the reference ligand. Four proteins obtained by this approach were produced for experimental evaluation. The X-ray structure of one of these proteins was determined to check for similarity of the transferred functional motif with the structure it adopts in the reference ligand. Three of these protein ligands bind the Kv1.2 channel with inhibition constants of 0.5, 1.5, and 1.6 microM. Several mutants of these designed protein ligands gave binding results consistent with the presumed binding mode. These results show that protein ligands can be designed by transferring a binding motif on a protein host selected to reproduce the functional topology of this motif, irrespective to the secondary structure supporting the functional motif, if the host protein possesses steric and electrostatic properties compatible with the binding to the target. This result opens the way to the design of protein ligands by taking advantage of the considerable structural repertoire of the Protein Data Bank.
- Département d'Ingénierie et d'Etude des Protéines, DSV, CEA, CE-Saclay, 91191 Gif Sur Yvette Cedex, France.
Organizational Affiliation: