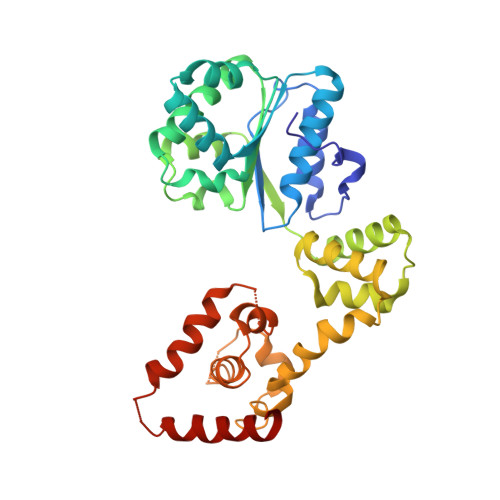
Structural basis for ATP-dependent DnaA assembly and replication-origin remodeling.
Erzberger, J.P., Mott, M.L., Berger, J.M.(2006) Nat Struct Mol Biol 13: 676-683
- PubMed: 16829961 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb1115
- Primary Citation Related Structures:
2HCB - PubMed Abstract:
In bacteria, the initiation of replication is controlled by DnaA, a member of the ATPases associated with various cellular activities (AAA+) protein superfamily. ATP binding allows DnaA to transition from a monomeric state into a large oligomeric complex that remodels replication origins, triggers duplex melting and facilitates replisome assembly. The crystal structure of AMP-PCP-bound DnaA reveals a right-handed superhelix defined by specific protein-ATP interactions. The observed quaternary structure of DnaA, along with topology footprint assays, indicates that a right-handed DNA wrap is formed around the initiation nucleoprotein complex. This model clarifies how DnaA engages and unwinds bacterial origins and suggests that additional, regulatory AAA+ proteins engage DnaA at filament ends. Eukaryotic and archaeal initiators also have the structural elements that promote open-helix formation, indicating that a spiral, open-ring AAA+ assembly forms the core element of initiators in all domains of life.
- Division of Biochemistry and Molecular Biology, Molecular and Cell Biology Department, 327B Hildebrand Hall #3206, University of California, Berkeley, California 94720, USA.
Organizational Affiliation: