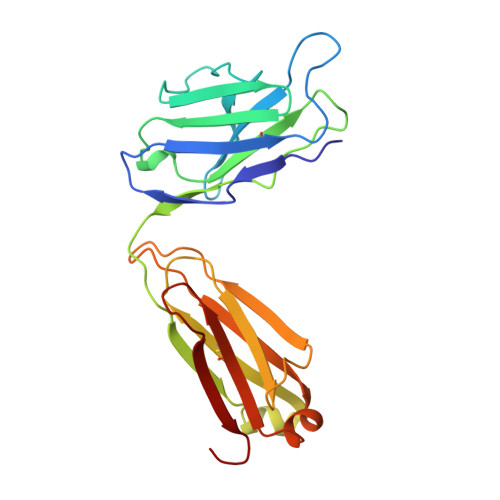
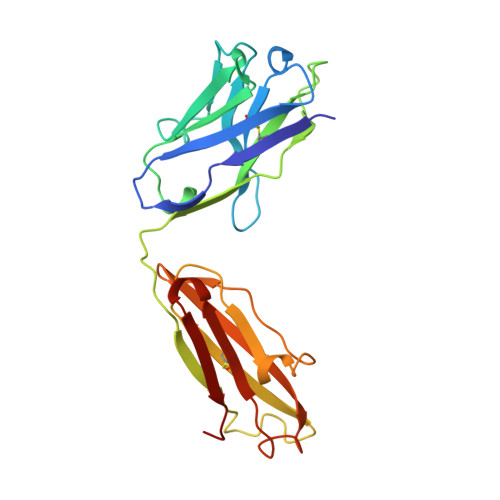
The three-dimensional structures of a polysaccharide binding antibody to Cryptococcus neoformans and its complex with a peptide from a phage display library: implications for the identification of peptide mimotopes.
Young, A.C., Valadon, P., Casadevall, A., Scharff, M.D., Sacchettini, J.C.(1997) J Mol Biology 274: 622-634
- PubMed: 9417940
- DOI: https://doi.org/10.1006/jmbi.1997.1407
- Primary Citation of Related Structures:
2H1P - PubMed Abstract:
The three-dimensional structure of 2H1, a protective monoclonal antibody to Cryptococcus neoformans, has been solved at 2.4 A resolution, in both its unbound form and in complex with the 12 amino acid residue peptide PA1 (GLQYTPSWMLVG). PA1 was previously identified as a potential mimotope of the cryptococcal capsular polysaccharide by screening of a phage display peptide library. Peptide binding is associated with only minor rearrangements of some side-chains and a small shift in the H2 loop of the antibody. The peptide assumes a tightly coiled conformation consisting of one inverse gamma-turn and one type II beta-turn that serves to place the entire peptide motif, consisting of ThrP5, ProP6, TrpP8, MetP9 and LeuP10, into a depression in the antibody combining site. A small number of H-bonds between peptide and antibody contribute to the affinity and specificity. Poor steric complementarity between PA1 and the antibody heavy chain along with the fact that the majority of the interactions between 2H1 and PA1 involve van der Waals interactions with the light chain may explain why this peptide acts as only a partial mimotope of the capsular polysaccharide epitope.
- Department of Microbiology, Albert Einstein College of Medicine, Bronx, NY 10461, USA.
Organizational Affiliation: