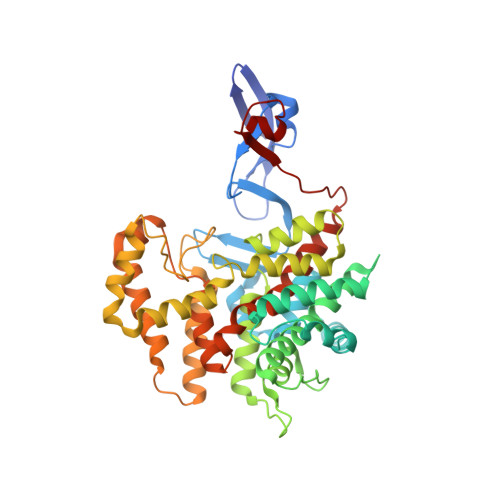
Structure of a NADH-insensitive hexameric citrate synthase that resists acid inactivation.
Francois, J.A., Starks, C.M., Sivanuntakorn, S., Jiang, H., Ransome, A.E., Nam, J.W., Constantine, C.Z., Kappock, T.J.(2006) Biochemistry 45: 13487-13499
- PubMed: 17087502
- DOI: https://doi.org/10.1021/bi061083k
- Primary Citation Related Structures:
2H12 - PubMed Abstract:
Acetobacter aceti converts ethanol to acetic acid, and strains highly resistant to both are used to make vinegar. A. aceti survives acetic acid exposure by tolerating cytoplasmic acidification, which implies an unusual adaptation of cytoplasmic components to acidic conditions. A. aceti citrate synthase (AaCS), a hexameric type II citrate synthase, is required for acetic acid resistance and, therefore, would be expected to function at low pH. Recombinant AaCS has intrinsic acid stability that may be a consequence of strong selective pressure to function at low pH, and unexpectedly high thermal stability for a protein that has evolved to function at approximately 30 degrees C. The crystal structure of AaCS, complexed with oxaloacetate (OAA) and the inhibitor carboxymethyldethia-coenzyme A (CMX), was determined to 1.85 A resolution using protein purified by a tandem affinity purification procedure. This is the first crystal structure of a "closed" type II CS, and its active site residues interact with OAA and CMX in the same manner observed in the corresponding type I chicken CS.OAA.CMX complex. While AaCS is not regulated by NADH, it retains many of the residues used by Escherichia coli CS (EcCS) for NADH binding. The surface of AaCS is abundantly decorated with basic side chains and has many fewer uncompensated acidic charges than EcCS; this constellation of charged residues is stable in varied pH environments and may be advantageous in the A. aceti cytoplasm.
- Department of Chemistry, Washington University, St. Louis, Missouri 63130-4899, USA.
Organizational Affiliation: