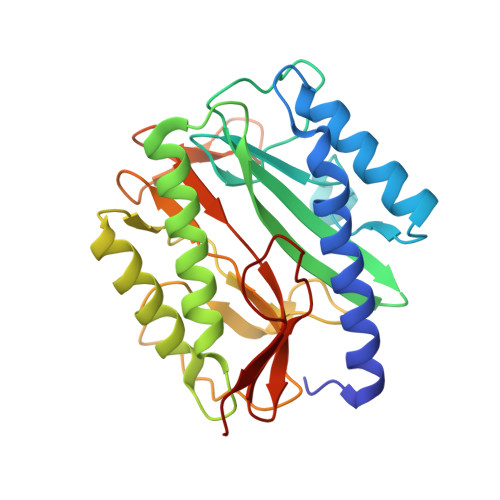
Structural basis of catalysis by monometalated methionine aminopeptidase.
Ye, Q.Z., Xie, S.X., Ma, Z.Q., Huang, M., Hanzlik, R.P.(2006) Proc Natl Acad Sci U S A 103: 9470-9475
- PubMed: 16769889 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0602433103
- Primary Citation Related Structures:
2GTX, 2GU4, 2GU5, 2GU6, 2GU7 - PubMed Abstract:
Methionine aminopeptidase (MetAP) removes the amino-terminal methionine residue from newly synthesized proteins, and it is a target for the development of antibacterial and anticancer agents. Available x-ray structures of MetAP, as well as other metalloaminopeptidases, show an active site containing two adjacent divalent metal ions bridged by a water molecule or hydroxide ion. The predominance of dimetalated structures leads naturally to proposed mechanisms of catalysis involving both metal ions. However, kinetic studies indicate that in many cases, only a single metal ion is required for full activity. By limiting the amount of metal ion present during crystal growth, we have now obtained a crystal structure for a complex of Escherichia coli MetAP with norleucine phosphonate, a transition-state analog, and only a single Mn(II) ion bound at the active site in the position designated M1, and three related structures of the same complex that show the transition from the mono-Mn(II) form to the di-Mn(II) form. An unliganded structure was also solved. In view of the full kinetic competence of the monometalated MetAP, the much weaker binding constant for occupancy of the M2 site compared with the M1 site, and the newly determined structures, we propose a revised mechanism of peptide bond hydrolysis by E. coli MetAP. We also suggest that the crystallization of dimetalated forms of metallohydrolases may, in some cases, be a misleading experimental artifact, and caution must be taken when structures are generated to aid in elucidation of reaction mechanisms or to support structure-aided drug design efforts.
- High Throughput Screening Laboratory and Department of Medicinal Chemistry, University of Kansas, 1501 Wakarusa Drive, Lawrence, KS 66045, USA. qye@ku.edu
Organizational Affiliation: