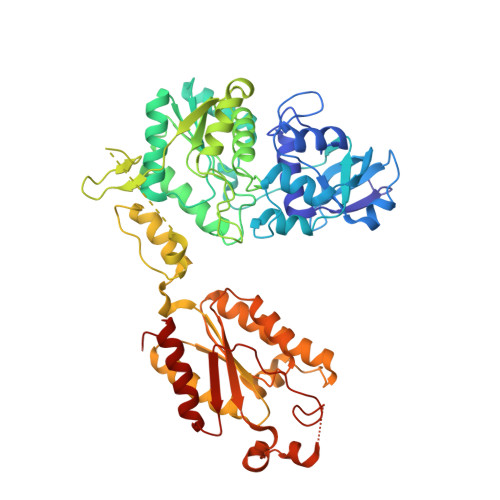
Crystal structure of the bifunctional ATP sulfurylase-APS kinase from the chemolithotrophic thermophile Aquifex aeolicus.
Yu, Z., Lansdon, E.B., Segel, I.H., Fisher, A.J.(2007) J Mol Biology 365: 732-743
- PubMed: 17095009
- DOI: https://doi.org/10.1016/j.jmb.2006.10.035
- Primary Citation Related Structures:
2GKS - PubMed Abstract:
The thermophilic chemolithotroph, Aquifex aeolicus, expresses a gene product that exhibits both ATP sulfurylase and adenosine-5'-phosphosulfate (APS) kinase activities. These enzymes are usually segregated on two separate proteins in most bacteria, fungi, and plants. The domain arrangement in the Aquifex enzyme is reminiscent of the fungal ATP sulfurylase, which contains a C-terminal domain that is homologous to APS kinase yet displays no kinase activity. Rather, in the fungal enzyme, the motif serves as a sulfurylase regulatory domain that binds the allosteric effector 3'-phosphoadenosine-5'-phosphosulfate (PAPS), the product of true APS kinase. Therefore, the Aquifex enzyme may represent an ancestral homolog of a primitive bifunctional enzyme, from which the fungal ATP sulfurylase may have evolved. In heterotrophic sulfur-assimilating organisms such as fungi, ATP sulfurylase catalyzes the first committed step in sulfate assimilation to produce APS, which is subsequently metabolized to generate all sulfur-containing biomolecules. In contrast, ATP sulfurylase in sulfur chemolithotrophs catalyzes the reverse reaction to produce ATP and sulfate from APS and pyrophosphate. Here, the 2.3 A resolution X-ray crystal structure of Aquifex ATP sulfurylase-APS kinase bifunctional enzyme is presented. The protein dimerizes through its APS kinase domain and contains ADP bound in all four active sites. Comparison of the Aquifex ATP sulfurylase active site with those from sulfate assimilators reveals similar dispositions of the bound nucleotide and nearby residues. This suggests that minor perturbations are responsible for optimizing the kinetic properties for the physiologically relevant direction. The APS kinase active-site lid adopts two distinct conformations, where one conformation is distorted by crystal contacts. Additionally, a disulfide bond is observed in one ATP-binding P-loop of the APS kinase active site. This linkage accounts for the low kinase activity of the enzyme under oxidizing conditions. The thermal stability of the Aquifex enzyme can be explained by the 43% decreased cavity volume found within the protein core.
- Department of Chemistry, University of California-Davis, One Shields Avenue, Davis, CA 95616, USA.
Organizational Affiliation: