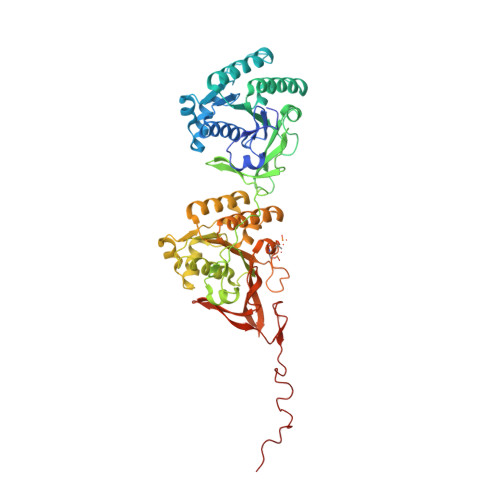
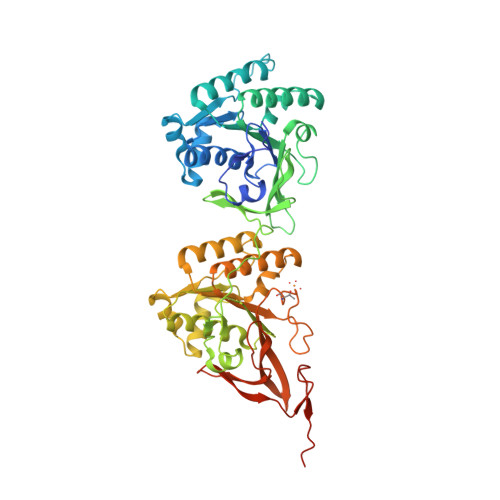
Analysis of KaiA-KaiC protein interactions in the cyano-bacterial circadian clock using hybrid structural methods.
Pattanayek, R., Williams, D.R., Pattanayek, S., Xu, Y., Mori, T., Johnson, C.H., Stewart, P.L., Egli, M.(2006) EMBO J 25: 2017-2028
- PubMed: 16628225 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601086
- Primary Citation Related Structures:
2GBL - PubMed Abstract:
The cyanobacterial circadian clock can be reconstituted in vitro by mixing recombinant KaiA, KaiB and KaiC proteins with ATP, producing KaiC phosphorylation and dephosphorylation cycles that have a regular rhythm with a ca. 24-h period and are temperature-compensated. KaiA and KaiB are modulators of KaiC phosphorylation, whereby KaiB antagonizes KaiA's action. Here, we present a complete crystallographic model of the Synechococcus elongatus KaiC hexamer that includes previously unresolved portions of the C-terminal regions, and a negative-stain electron microscopy study of S. elongatus and Thermosynechococcus elongatus BP-1 KaiA-KaiC complexes. Site-directed mutagenesis in combination with EM reveals that KaiA binds exclusively to the CII half of the KaiC hexamer. The EM-based model of the KaiA-KaiC complex reveals protein-protein interactions at two sites: the known interaction of the flexible C-terminal KaiC peptide with KaiA, and a second postulated interaction between the apical region of KaiA and the ATP binding cleft on KaiC. This model brings KaiA mutation sites that alter clock period or abolish rhythmicity into contact with KaiC and suggests how KaiA might regulate KaiC phosphorylation.
- Department of Biochemistry, School of Medicine, Vanderbilt University, Nashville, TN 37232, USA.
Organizational Affiliation: