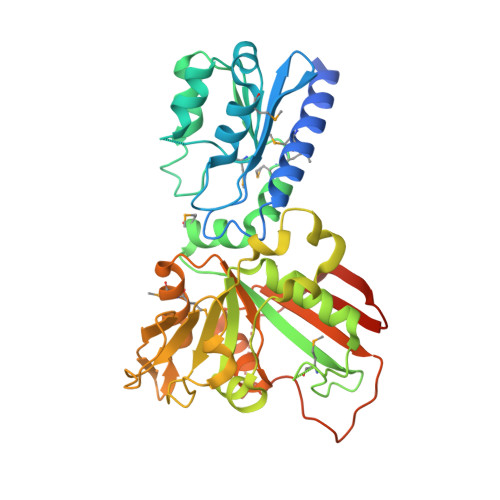
The Crystal Structure of the Bifunctional Deaminase/Reductase RibD of the Riboflavin Biosynthetic Pathway in Escherichia coli: Implications for the Reductive Mechanism.
Stenmark, P., Moche, M., Gurmu, D., Nordlund, P.(2007) J Mol Biology 373: 48-64
- PubMed: 17765262 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.12.009
- Primary Citation Related Structures:
2G6V, 2O7P, 2OBC - PubMed Abstract:
We have determined the crystal structure of the bi-functional deaminase/reductase enzyme from Escherichia coli (EcRibD) that catalyzes two consecutive reactions during riboflavin biosynthesis. The polypeptide chain of EcRibD is folded into two domains where the 3D structure of the N-terminal domain (1-145) is similar to cytosine deaminase and the C-terminal domain (146-367) is similar to dihydrofolate reductase. We showed that EcRibD is dimeric and compared our structure to tetrameric RibG, an ortholog from Bacillus subtilis (BsRibG). We have also determined the structure of EcRibD in two binary complexes with the oxidized cofactor (NADP(+)) and with the substrate analogue ribose-5-phosphate (RP5) and superposed these two in order to mimic the ternary complex. Based on this superposition we propose that the invariant Asp200 initiates the reductive reaction by abstracting a proton from the bound substrate and that the pro-R proton from C4 of the cofactor is transferred to C1 of the substrate. A highly flexible loop is found in the reductase active site (159-173) that appears to control cofactor and substrate binding to the reductase active site and was therefore compared to the corresponding Met20 loop of E. coli dihydrofolate reductase (EcDHFR). Lys152, identified by comparing substrate analogue (RP5) coordination in the reductase active site of EcRibD with the homologous reductase from Methanocaldococcus jannaschii (MjaRED), is invariant among bacterial RibD enzymes and could contribute to the various pathways taken during riboflavin biosynthesis in bacteria and yeast.
- Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-171 77 Stockholm, Sweden.
Organizational Affiliation: