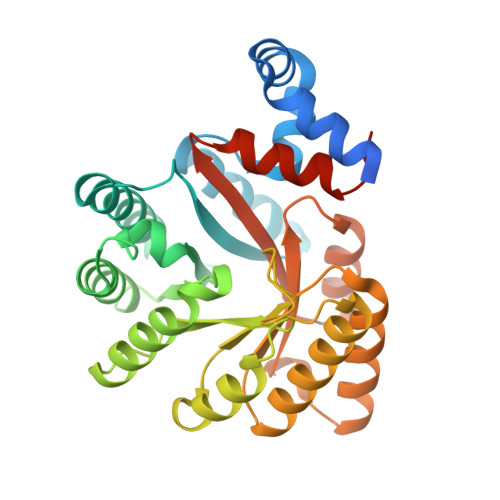
Structure and Kinetics of Monofunctional Proline Dehydrogenase from Thermus thermophilus.
White, T.A., Krishnan, N., Becker, D.F., Tanner, J.J.(2007) J Biological Chem 282: 14316-14327
- PubMed: 17344208
- DOI: https://doi.org/10.1074/jbc.M700912200
- Primary Citation Related Structures:
2G37 - PubMed Abstract:
Proline dehydrogenase (PRODH) and Delta(1)-pyrroline-5-carboxylate dehydrogenase (P5CDH) catalyze the two-step oxidation of proline to glutamate. They are distinct monofunctional enzymes in all eukaryotes and some bacteria but are fused into bifunctional enzymes known as proline utilization A (PutA) in other bacteria. Here we report the first structure and biochemical data for a monofunctional PRODH. The 2.0-A resolution structure of Thermus thermophilus PRODH reveals a distorted (betaalpha)(8) barrel catalytic core domain and a hydrophobic alpha-helical domain located above the carboxyl-terminal ends of the strands of the barrel. Although the catalytic core is similar to that of the PutA PRODH domain, the FAD conformation of T. thermophilus PRODH is remarkably different and likely reflects unique requirements for membrane association and communication with P5CDH. Also, the FAD of T. thermophilus PRODH is highly solvent-exposed compared with PutA due to a 4-A shift of helix 8. Structure-based sequence analysis of the PutA/PRODH family led us to identify nine conserved motifs involved in cofactor and substrate recognition. Biochemical studies show that the midpoint potential of the FAD is -75 mV and the kinetic parameters for proline are K(m) = 27 mm and k(cat) = 13 s(-1). 3,4-Dehydro-l-proline was found to be an efficient substrate, and l-tetrahydro-2-furoic acid is a competitive inhibitor (K(I) = 1.0 mm). Finally, we demonstrate that T. thermophilus PRODH reacts with O(2) producing superoxide. This is significant because superoxide production underlies the role of human PRODH in p53-mediated apoptosis, implying commonalities between eukaryotic and bacterial monofunctional PRODHs.
- Department of Biochemistry, University of Missouri-Columbia, Columbia, MO 65211, USA.
Organizational Affiliation: