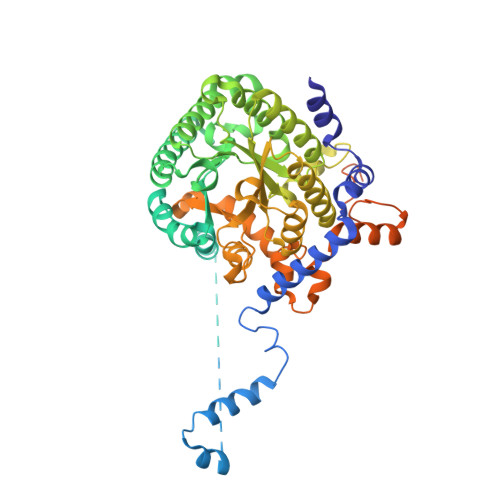
Redox-induced changes in flavin structure and roles of flavin N(5) and the ribityl 2'-OH group in regulating PutA--membrane binding.
Zhang, W., Zhang, M., Zhu, W., Zhou, Y., Wanduragala, S., Rewinkel, D., Tanner, J.J., Becker, D.F.(2007) Biochemistry 46: 483-491
- PubMed: 17209558 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi061935g
- Primary Citation Related Structures:
2FZM - PubMed Abstract:
PutA is a novel flavoprotein in Escherichia coli that switches from a transcriptional repressor to a membrane-bound proline catabolic enzyme. Previous crystallographic studies of the PutA proline dehydrogenase (PRODH) domain under oxidizing conditions revealed that FAD N(5) and the ribityl 2'-OH group form hydrogen bonds with Arg431 and Arg556, respectively. Here we identify molecular interactions in the PutA PRODH active site that underlie redox-dependent functional switching of PutA. We report that reduction of the PRODH domain induces major structural changes in the FAD cofactor, including a 22 degrees bend of the isoalloxazine ring along the N(5)-N(10) axis, crankshaft rotation of the upper part of the ribityl chain, and formation of a new hydrogen bond network involving the ribityl 2'-OH group, FAD N(1), and Gly435. The roles of the FAD 2'-OH group and the FAD N(5)-Arg431 hydrogen bond pair in regulating redox-dependent PutA-membrane associations were tested using FAD analogues and site-directed mutagenesis. Kinetic membrane binding measurements and cell-based reporter gene assays of modified PutA proteins show that disrupting the FAD N(5)-Arg431 interaction impairs the reductive activation of PutA-membrane binding. We also show that the FAD 2'-OH group acts as a redox-sensitive toggle switch that controls PutA-membrane binding. These results illustrate a new versatility of the ribityl chain in flavoprotein mechanisms.
- Department of Biochemistry, University of Nebraska, Lincoln, Nebraska 68588, USA.
Organizational Affiliation: